Call us
301-363-4651 (Available 9 a.m. to 5 p.m. CST from Monday to Friday)
| Code | CSB-E17414r |
| Size | 96T,5×96T,10×96T |
| Price | Request a Quote |
| Trial Size |
24T ELISA Kit Trial Size (Only USD$150/ kit) * Sample kit cost can be deducted as a $30 credit for each 96-assay kit of the same analyte and brand you subsequently purchase within six months until depleted. More details >> Interested in a trial size? Please leave a message below.
|
| Have Questions? | Leave a Message or Start an on-line Chat |
| Intra-assay Precision (Precision within an assay): CV%<8% | ||||||
| Three samples of known concentration were tested twenty times on one plate to assess. | ||||||
| Inter-assay Precision (Precision between assays): CV%<10% | ||||||
| Three samples of known concentration were tested in twenty assays to assess. | ||||||
| To assess the linearity of the assay, samples were spiked with high concentrations of rat SREBP-1C in various matrices and diluted with the Sample Diluent to produce samples with values within the dynamic range of the assay. | ||||||
| Sample | Serum(n=4) | |||||
| 1:5 | Average % | 90 | ||||
| Range % | 86-96 | |||||
| 1:10 | Average % | 101 | ||||
| Range % | 95-108 | |||||
| 1:20 | Average % | 95 | ||||
| Range % | 90-100 | |||||
| 1:40 | Average % | 92 | ||||
| Range % | 87-96 | |||||
| The recovery of rat SREBP-1C spiked to levels throughout the range of the assay in various matrices was evaluated. Samples were diluted prior to assay as directed in the Sample Preparation section. | ||||||
| Sample Type | Average % Recovery | Range | ||||
| Serum (n=5) | 93 | 90-96 | ||||
| EDTA plasma (n=4) | 96 | 90-102 | ||||
| These standard curves are provided for demonstration only. A standard curve should be generated for each set of samples assayed. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
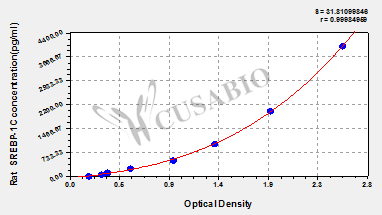
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
This Rat SREBP-1C ELISA Kit was designed for the quantitative measurement of Rat SREBP-1C protein in serum, plasma, tissue homogenates. It is a Sandwich ELISA kit, its detection range is 62.5 pg/mL-4000 pg/mL and the sensitivity is 15.6 pg/mL .
There are currently no reviews for this product.