Modified Histone Antibodies
Epigenetics is the study of heritable phenotype changes in gene expression that modify DNA, RNA, and protein but do not involve alterations of the nucleotide sequence. Posttranslational modifications (PTMs) are the most important method to research epigenetics that apply to proteins. PTMs are marks that provide an extensive regulatory mechanism for cells to signal which genes to turn off and on. Many types and families of proteins are subject to PTMs, but one of the most highly decorated is the histone family of proteins.
Histone packages genomic DNA into nucleosomes, which allows about 2 meters of DNA to be placed in the nucleus of a cell. The nucleosome contains two subunits, each consisting of histone H2A, H2B, H3, and H4. In addition, there is histone H1, commonly referred to as a junctional histone. The most prevalent PTMs found on histones have a crucial role in gene expression, including methylation, acetylation, phosphorylation, and ubiquitination. These markers serve as signals for opening and compressing chromatin, as well as recruitment factors that promote and antagonize transcription. The modifications of histone on below image can regulate gene expression, which in turn affects chromosome function.
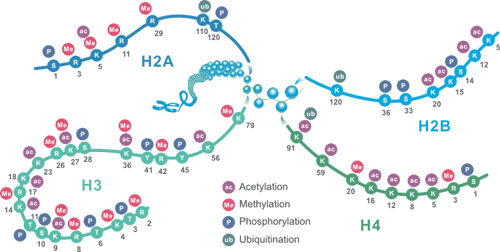
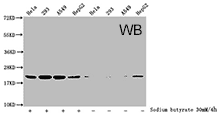
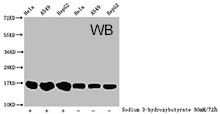
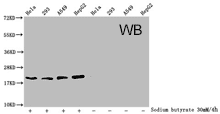
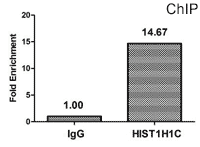
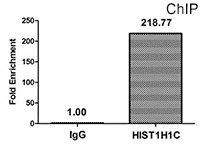
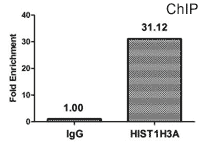
CUSABIO provides highly specific antibodies to histone modifications, including histone methylation antibodies, histone phosphorylation antibodies, histone acetylation antibodies, histone ubiquitylation antibodies, etc. They have been validated by ChIP, WB, IHC, IP, etc., and can be used to detect modified histones in specific gene regions.
Product Features
- Highly specific recognition of modified histone
- Well validation of ChIP application
CUSABIO Modified Histone Antibodies catalog
Here is the catalog of some Modified Histone Antibodies.
Useful Resource
What is histone modification?
Histone modification are covalent and enzymatic post-translational modifications (PTMs) of histones by various modifying enzymes. The PTMs made to histones can regulate gene transcription through the alteration of chromatin structure by introducing different histone modifiers to histone amino acid residues.
What is the purpose of histone modification?
Histone modifications mediate various biological processes such as transcriptional activation or inhibition, chromosome packaging, heterochromatin formation, nucleosome assembly and disassembly, and DNA damage repair. Numerous different diseases such as cancer, autoimmune disorders and inflammatory and neurological diseases have been associated with abnormal histone modifications. These histone modifications all together make up what is known as the histone code, which exhibits the transcriptional state of the local genomic region including promoters, enhancers, and other gene regulatory elements. Therefore, quantitative detection of histone modifications would uncover novel epigenetic insight into cellular processes and disease states, which is beneficial to the development of drugs that are targeted histone modification.
How many histone modifications are there? And what are they?
There are at least 15 different known modifications associated with the histones. Among these histone modifications, acetylation, methylation, phosphorylation, and ubiquitylation are the most well-studied, while sumoylation, isomerization, ADP-ribosylation, and biotinylation, etc. have yet not been thoroughly investigated.
- Histone methylation is the modification of certain amino acids in a histone protein by the addition of methyl groups. It causes transcription repression or activation, depending on the target sites. Histone methyltransferases (HMTs) control or regulate DNA methylation through chromatin-dependent transcription repression or activation. Methylation and demethylation of histones turn genes on and off in DNA.
- Histone phosphorylation has been implicated in numerous processes such as transcription, DNA repair, cell cycle progression and apoptosis, and chromosome condensation. The best-known function of histone phosphorylation takes place during cellular response to DNA damage, when phosphorylated histone H2A(X) demarcates large chromatin domains around the site of DNA breakage.
- Histone acetylation is triggered by the enzymatic addition of an acetyl group (COCH3) from acetyl coenzyme A. The process of histone acetylation is closely involved in the regulation of many cellular processes, including chromatin dynamics and transcription, gene silencing, cell cycle progression, apoptosis, differentiation, DNA replication, DNA repair, nuclear import, and neuronal repression.
- Histone succinylation is a posttranslational modification where a succinyl group (-CO-CH2-CH2-COOH) is added to a lysine residue of histone. The potential role of succinylation is under investigation, but as addition of succinyl group changes lysine's charge from +1 to −1 (at physiological pH) and introduces a relatively large structural moiety (100 Da), bigger than acetylation (42 Da) or methylation (14 Da), it is expected to lead to more significant changes in protein structure and function.
- Histone 2-hydroxyisobutyrylation is a new type of histone mark and identified at 63 human and mouse histone Khib sites, including 27 unique lysine sites that are not known to be modified by lysine acetylation (Kac) and lysine crotonylation (Kcr). This histone mark was initially identified by MS and then validated by chemical and biochemical methods.
- Histone biotinylation is a relatively new field of research, evidence of biological roles for biotinylation of histones is scarce. Recently, emerging evidence has suggested that biotinylation of histones increases in response to cell proliferation in human lymphocytes. Moreover, biotinylation of histones might play a role in the cellular response to DNA damage.
- Histone formylation is one of many post-translational modifications, which been shown to modulate chromatin conformations and gene activation. Formylation has been identified on the Nε of lysine residues in histones and proteins. This modification has been observed in linker histones and high mobility group proteins, it is highly abundant and it is believed to have a role in the epigenetics of chromatin function.
- Histone crotonylation is a newly identified histone modification that is associated with active transcription in mammalian cells. The story of crotonylation began in 2011 with a paper from the lab of Yingming Zhao when they discovered it as a novel histone modification on lysine residues. The Zhao lab showed histone crotonylation correlated well with gene expression and marked promoters and enhancers just like its close cousin, acetylation.
- Histone hydroxylation is one of many post-translational modification that involves conversion of a CH group into a COH group. Hydroxylation is an oxidative process. Several studies have revealed that histone 5-hydroxylation may have important roles in epigenetic regulation of gene transcription or chromosomal rearrangement.
- Histone O-GlcNAcylation is a posttranslational modification. Currently, three sites of O-GlcNAcylation were mapped by MS/MS on histone H2A Thr101, histone H2B Ser36, and histone H4 Ser47, respectively.
- Histone malonylation is a new types of histone PTMs. The study from Zhongyu Xie team has demonstrated that lysine malonylation induce even more significant structural changes than lysine acetylation by changing a positively charged residue to a negative charge.
- Histone glutarylation is one of many posttranslational modification, which usually occurs on lysine. Kristin A. Anderson et al. identified and comprehensively validated lysine glutarylation as an evolutionarily conserved PTM, and impacted metabolic processes and other mitochondrial functions.
- Histone β‑hydroxybutyrylation is a new type of histone modification, which usually occurs on lysine. According to the study of Zhongyu Xie team, this modification is induced significantly during prolonged fasting in mouse liver and is associated with genes up-regulated in starvation-responsive metabolic pathways.