Full Product Name
Rabbit anti-Homo sapiens (Human) H3F3A Polyclonal antibody
Alternative Names
H3 histone family 3A antibody; H3 histone family 3B antibody; H3 histone; family 3B (H3.3B) antibody; H3.3 antibody; H3.3A antibody; H3.3B antibody; H33_HUMAN antibody; H3F3 antibody; H3F3A antibody; H3f3b antibody; Histone H3.3 antibody; Histone H3.3Q antibody; Histone H3.A antibody; Histone H3.B antibody; MGC87782 antibody; MGC87783 antibody
Immunogen
Peptide sequence around site of Di-methyl-Lys (79) derived from Human Histone H3.3
Immunogen Species
Homo sapiens (Human)
Purification Method
Antigen Affinity Purified
Concentration
It differs from different batches. Please contact us to confirm it.
Buffer
Preservative: 0.03% Proclin 300
Constituents: 50% Glycerol, 0.01M PBS, pH 7.4
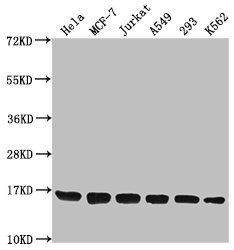
Tested Applications
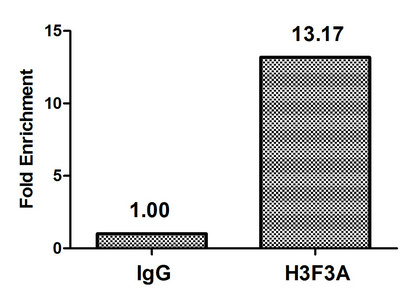
ELISA, WB, ChIP
Recommended Dilution
| Application |
Recommended Dilution |
| WB |
1:500-1:2000 |
Storage
Upon receipt, store at -20°C or -80°C. Avoid repeated freeze.
Lead Time
Basically, we can dispatch the products out in 1-3 working days after receiving your orders. Delivery time maybe differs from different purchasing way or location, please kindly consult your local distributors for specific delivery time.
Usage
For Research Use Only. Not for use in diagnostic or therapeutic procedures.