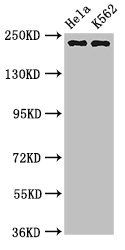
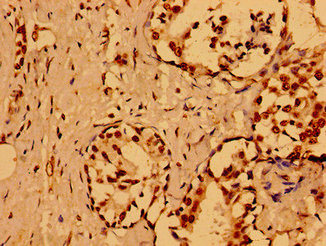
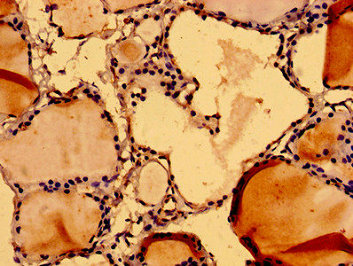
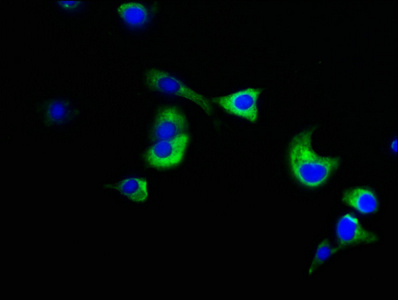
Function
Microtubule (MT)-binding protein that plays a role in the formation and maintenance of the spindle poles and the alignement and the segregation of chromosomes during mitotic cell division. Functions to tether the minus ends of MTs at the spindle poles, which is critical for the establishment and maintenance of the spindle poles. Plays a role in the establishment of the mitotic spindle orientation during metaphase and elongation during anaphase in a dynein-dynactin-dependent manner. In metaphase, part of a ternary complex composed of GPSM2 and G(i) alpha proteins, that regulates the recruitment and anchorage of the dynein-dynactin complex in the mitotic cell cortex regions situated above the two spindle poles, and hence regulates the correct oritentation of the mitotic spindle. During anaphase, mediates the recruitment and accumulation of the dynein-dynactin complex at the cell membrane of the polar cortical region through direct association with phosphatidylinositol 4,5-bisphosphate (PI(4,5)P2), and hence participates in the regulation of the spindle elongation and chromosome segregation. Binds also to other polyanionic phosphoinositides, such as phosphatidylinositol 3-phosphate (PIP), lysophosphatidic acid (LPA) and phosphatidylinositol triphosphate (PIP3), in vitro. Also required for proper orientation of the mitotic spindle during asymmetric cell divisions. Plays a role in mitotic MT aster assembly. Involved in anastral spindle assembly. Positively regulates TNKS protein localization to spindle poles in mitosis. Highly abundant component of the nuclear matrix where it may serve a non-mitotic structural role, occupies the majority of the nuclear volume. Required for epidermal differentiation and hair follicle morphogenesis.
Subcellular Location
Nucleus. Nucleus, nucleoplasm. Nucleus matrix. Chromosome. Cytoplasm, cytoskeleton. Cytoplasm, cytoskeleton, microtubule organizing center, centrosome. Cytoplasm, cytoskeleton, spindle pole. Cytoplasm, cell cortex. Cell membrane; Lipid-anchor; Cytoplasmic side. Lateral cell membrane.; [Isoform 3]: Cytoplasm, cytosol. Cytoplasm, cytoskeleton, microtubule organizing center, centrosome. Cytoplasm, cytoskeleton, spindle pole.; [Isoform 4]: Cytoplasm, cytosol. Cytoplasm, cytoskeleton, microtubule organizing center, centrosome. Cytoplasm, cytoskeleton, spindle pole.