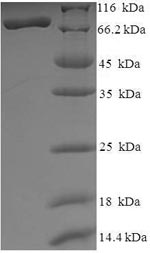
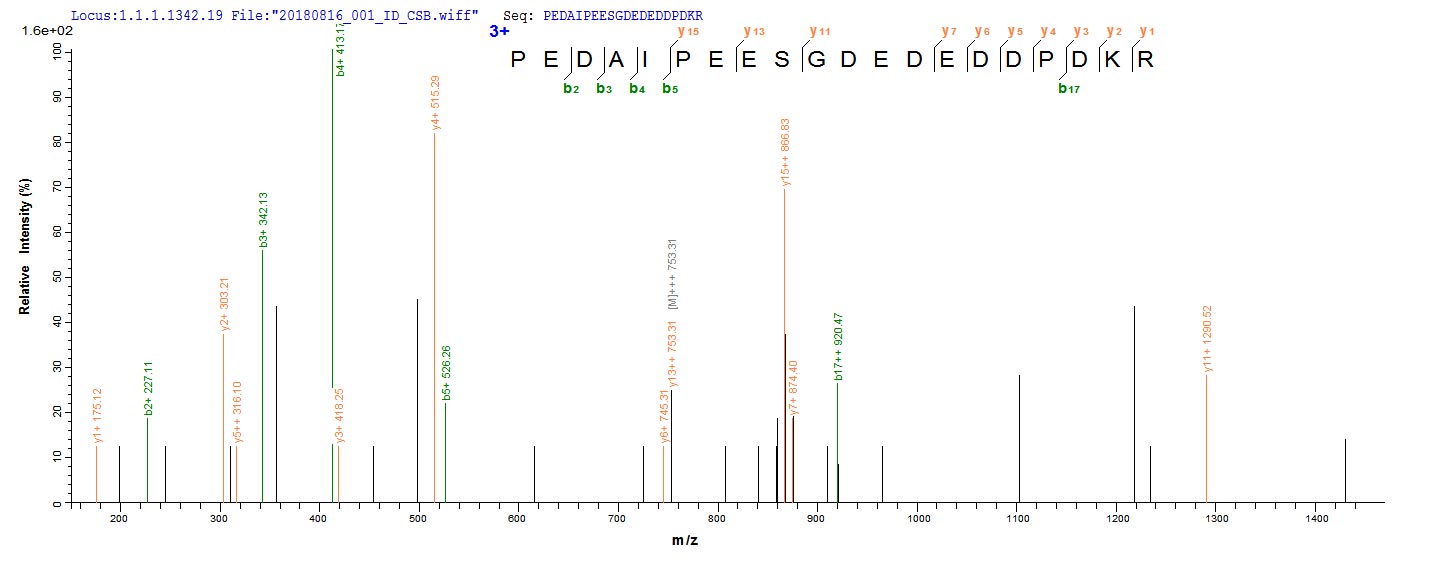
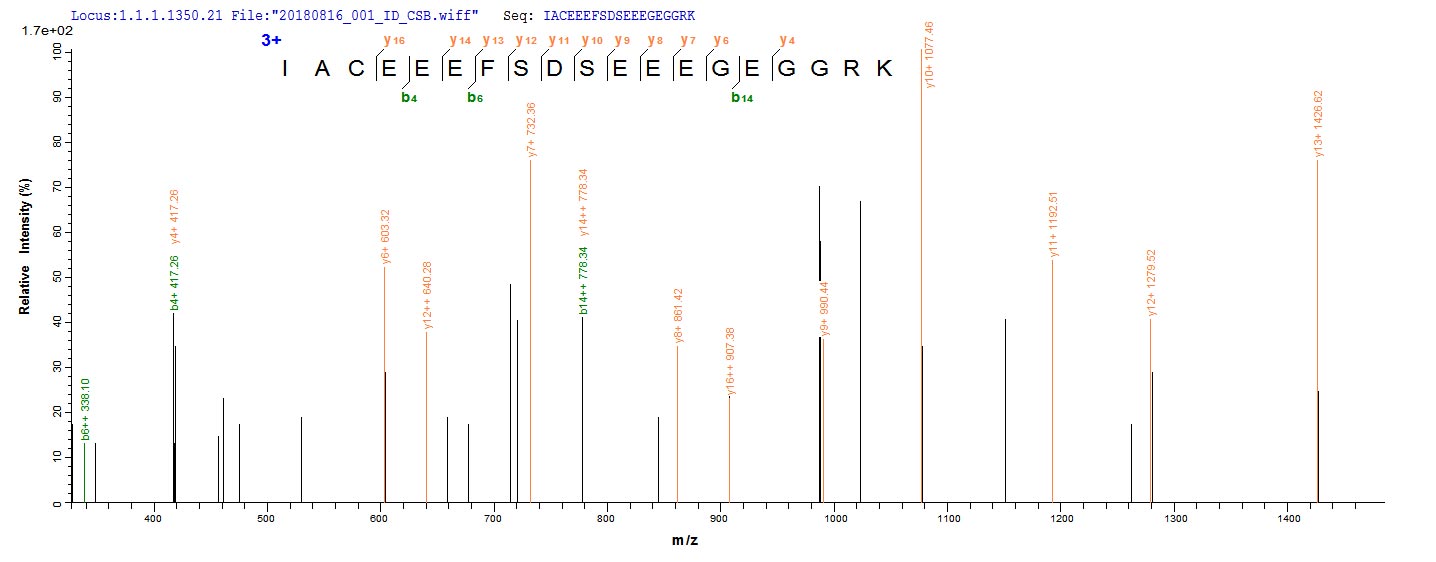
In the production of recombinant Human HDAC1 protein, the gene for HDAC1 (E.coli) was cloned into a vector and expressed as HDAC1 protein in E.coli. The plasmids with the copy of HDAC1, or the expression vector, were often used to enhance gene expression. Every step of production was undergone with a strict QC system. N-terminal 6xHis-SUMO tag was used in the process. The purity is 90% determined by SDS-PAGE.
HDAC1 belongs to the class IV family. HDAC1 is found in at least three evolutionally conserved, distinct protein complexes: the Sin3, the CoREST and the Mi-2/NuRD complexes. HDAC1 activity is modulated through various post translational modifications including phosphorylation, acetylation, ubiquitination, sumoylation, and nitrosylation. HDAC1 is phosphorylated at serine 421 and 423 by casein kinase II and dephosphorylated by mitogen-activated protein kinase phosphatase-3 (MKP-3). Mutagenesis analysis suggested that phosphorylation on these sites are important for histone deacetylase activity and for HDAC1 to interact with corepressor complexes. HDAC1 phosphorylation increases during the G1 phase and was significantly reduced during the late S/G2 phase. Clinically, it has been shown HDACi can reactivate fetal globin in adult erythroid cells, suggesting the role of HDAC1 in modulating key transcription factors for erythropoiesis and its clinical relevance.