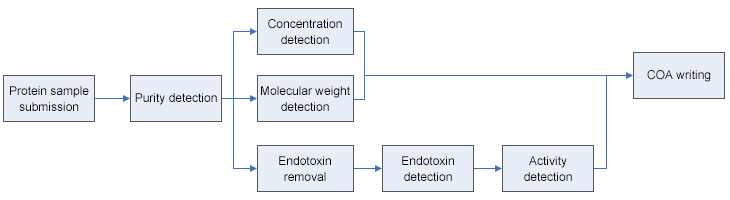
The quality of our proteins is the cornerstone of our ability to serve our clients. For this reason, we invest tremendous resources in developing proteins, validating their application, managing, controlling and improving quality. Each protein will undergo our standardized quality control process before shipped, which include concentration detection, purity detection, endotoxin removal, activity detection, et al.
1. Protein Concentration Detection
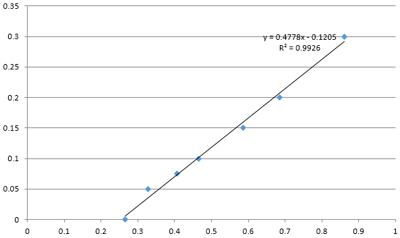
There are three methods of protein concentration detection: bradford method, BCA method, and A280 method. According to the cost of buffer, the bradford method is the most popular with laboratories, and the concentration detection range is 0.1-5 mg/ml. Bradford's standard curve is shown below:
2. Molecular Detection
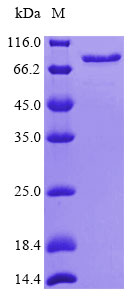
Protein molecular weight is detected by Coomassie brilliant blue staining combined with SDS-PAGE, and the molecular mass is consistent with the theoretical molecular mass predicted from the amino acid sequence. The theoretical molecular mass of Calreticulin protein is 73.3kDa. The test result is as follows:
3. Purity Detection
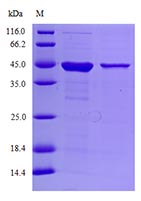
Protein purity was detected using Coomassie Brilliant Blue SDS-PAGE to isolate proteins. The bandscan software was used to analyze protein purity. The purity of the protein must reach 90%. The picture of Serine protease inhibitor A3N is as follows:
4. Endotoxin Detection
The endotoxin assay uses the LAL method of endotoxin levels below 1 EU/μg.
5. Biological Activity Detection
There are three methods of detecting protein activity: ELISA method, cell activity test, and enzyme activity detection method. Among them, the receptor protein mainly adopts ELISA method, the cytokine protein adopts cell activity detection method, and the enzyme uses enzyme activity detection method.
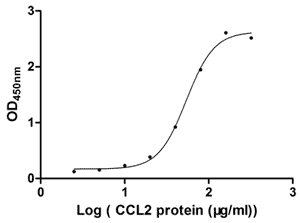
Measured by its binding ability in a functional ELISA. Immobilized ACKR1 at 1 μg/ml can bind human CCL2, the EC50 of human CCL2 protein is 48.64-60.24 μg/ml.
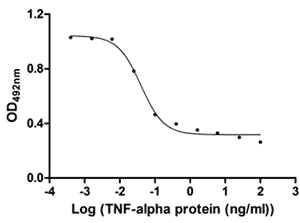
Measured in a cytotoxicity assay using L 929 mouse fibroblast cells in the presence of the metabolic inhibitor actinomycin D. The ED50 for this effect is 33.32-47.38 pg/ml
6. Sequence Accuracy
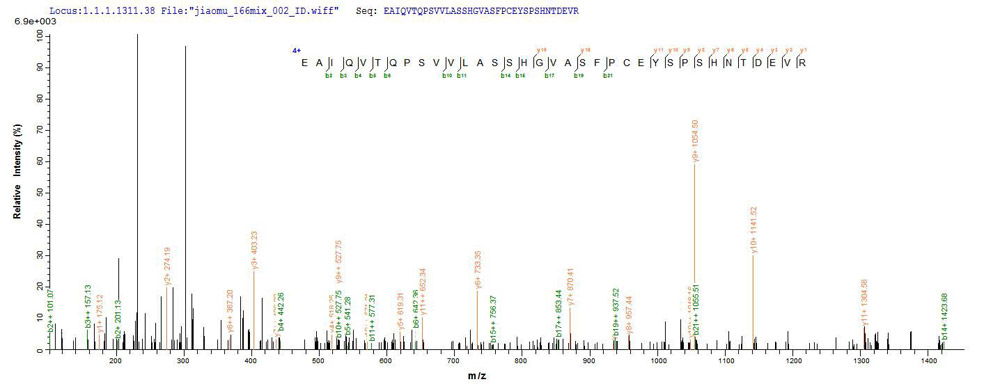
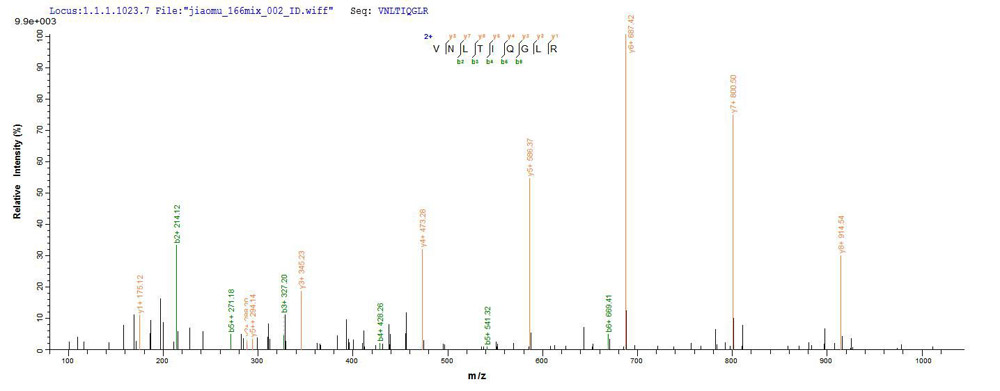
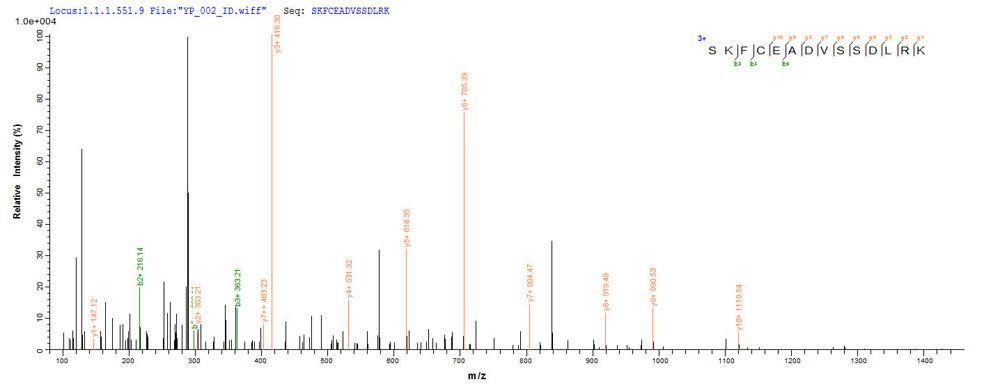
Protein Mass Spectrometry Identification is one of the core technologies in proteomics research. At present, Mass Spectrometry is the main method of identifying proteins. Mass spectrometry can be divided into two categories: MALDI-TOF and LC-MS / MS, among which, LC-MS / MS has a higher sensitivity and almost 100% reliability, which is the most commonly used method in the mainstream literature. Identifying a single protein from a mixture of protein samples is one of the issues in protein research. There are two common methods for separating protein mixtures: two-dimensional gel electrophoresis (2-DE) and polyacrylamide gel electrophoresis (SDS-PAGE, Sodium Dodecyl Sulfate Polyacrylamide Gel Electrophoresis).
Our protein gel strips and gel point identification have formed a set of mature processes, mainly including obtaining the strips and spots of the target protein by one-dimensional or two-dimensional electrophoresis, and optimizing the in-gel enzymolysis method serially, and then obtain protein fragmentation chip charge, peak maps and other related information by LC-MS / MS analysis. This method can significantly improve the success rate of mass spectrometry identification and the coverage of peptides in the in-gel digestion of protein.