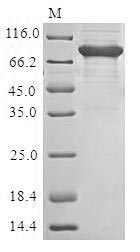
The Recombinant protein with an MBP-tag at N-terminal and a 6xHis-tag at the C-terminal corresponds to the amino acids 22-555 of Human Perforin-1/PRF1 full-length ORF. This Baculovirus-expressed PRF1 protein got purified by SDS-PAGE and reached up to 85% in purity. On a reducing SDS–PAGE gel, the PRF1 protein migrated to about 105kDa. Customers can get the protein faster because it has stock. The recombinant full-length of mature PRF1 protein may be used for antibody synthesis or in the immunological research related to PRF1.
PRF1 is a Ca2+-dependent glycoprotein mainly expressed by natural killer (NK)cells and CD8-positive T-cells. Polymerization of PRF1 forms channels on the target cell, thus allowing for the passive diffusion of granzymes into the cell. The cellular cytotoxicity mediated by PRF1 is responsible for killing virus-infected and neoplastic cells. Mutations in the PRF1 gene are associated with familial hemophagocytic lymphohistiocytosis (FHL) and various types of lymphomas.