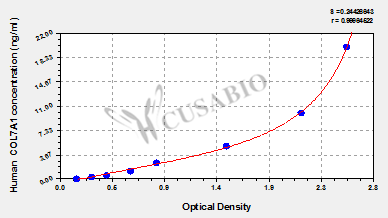
Per customers request, please provide the following information regarding Human Collagen alpha-1 (VII) chain, COL7A1 ELISA Kit (CSB-EL005754HU)
1)The active ingredient of the stop solution and its concentration ?
2)The type, concentration, and state (liquid/lyophilized) of the preservatives used ?
3) If the components are lyophilized, what is the concentration prior to reconstitution (i.e. mass of preservative used/ total mass of lyophilized vial contents)?
4)Expression host of the standards ?
5)Which components have H202 with the concentration of H202 in each, if any?
6) Are there any components that contain hydrogen peroxide & Urea hydrogen peroxide (Carbamide peroxide) in any of the components in this kit?
7) If yes, which components contain hydrogen peroxide & Urea hydrogen peroxide (Carbamide peroxide) and the concentration?