| Code | CSB-E12112h |
| Size | 96T,5×96T,10×96T |
| Price | Request a Quote or Start an on-line Chat |
| Trial Size |
24T ELISA Kit Trial Size (Only USD$150/ kit) * The sample kit cost can be deducted from your subsequent orders of 96T full size kits of the same analyte at 1/5 per kit, until depleted in 6 months. Apply now |
| Intra-assay Precision (Precision within an assay): CV%<8% | ||||||
| Three samples of known concentration were tested twenty times on one plate to assess. | ||||||
| Inter-assay Precision (Precision between assays): CV%<10% | ||||||
| Three samples of known concentration were tested in twenty assays to assess. | ||||||
| To assess the linearity of the assay, samples were spiked with high concentrations of human HIF-1α in various matrices and diluted with the Sample Diluent to produce samples with values within the dynamic range of the assay. | ||||||
| Sample | Serum(n=4) | |||||
| 1:1 | Average % | 90 | ||||
| Range % | 84-96 | |||||
| 1:2 | Average % | 96 | ||||
| Range % | 90-101 | |||||
| 1:4 | Average % | 99 | ||||
| Range % | 95-103 | |||||
| 1:8 | Average % | 94 | ||||
| Range % | 88-98 | |||||
| The recovery of human HIF-1α spiked to levels throughout the range of the assay in various matrices was evaluated. Samples were diluted prior to assay as directed in the Sample Preparation section. | ||||||
| Sample Type | Average % Recovery | Range | ||||
| Serum (n=5) | 95 | 91-98 | ||||
| EDTA plasma (n=4) | 96 | 92-99 | ||||
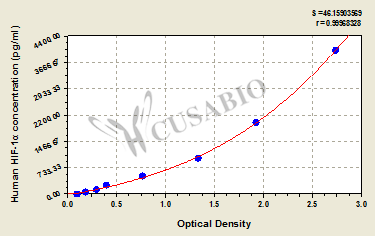
| These standard curves are provided for demonstration only. A standard curve should be generated for each set of samples assayed. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
The human HIF-1α ELISA Kit allows for the in vitro quantitative determination of HIF-1α concentrations in serum, plasma, or tissue homogenates. This kit exclusively recognizes human HIF-1α. HIF-1α is a nuclear transcription factor that has been identified as a biomarker of the hypoxia microenvironment in most types of solid tumors. Under hypoxia, HIF-1α becomes relatively stable and binds to HIF-1β to form the active HIF-1 heterodimer. Many studies have reported that HIF-1α is detected in the majority of solid tumors including breast cancer, gastric cancer, and cervical cancer, and is linked to cell proliferation, metabolism, immune responses, apoptosis, genomic instability, vascularization, invasion, and metastasis of tumor cells. HIF-1α is also associated with chemo-/radioresistance. Pharmacological HIF-1α suppression is thus required for reversing chemo-/radioresistance in tumors.
The detection mechanism of this kit is based on the Sandwich-ELISA technique. HIF-1α in the sample is bound to the capture antibody immobilized on the microtiter plate and then sandwiched with a Biotin-labeled HIF-1α antibody. The solution color develops into blue after the ordinal addition of HRP-avidin and TMB. The color development is terminated after adding the stop solution, and the color turns from blue to yellow. The color intensity is positively correlated with HIF-1α content in the sample. The kit has been quality-controlled with high sensitivity, strong specificity, good linearity, high precision, high recovery, and high lot-to-lot consistency.
There are currently no reviews for this product.
I wanna ask you a question please about the detection limit of the serum HIF-1α ELISA kit.
We are planning to measure serum HIF-1α levels in our study. But we are expecting low HIF-1α levels in our study subject. I mean, lower than 62.5 pg/mL, which is the lowest detection limit of the CUSABİO kit.
So, we will not able to use your HIF-1α kits, or maybe you can offer us another solution?
For the parameters of the kit CSB-E12112h, please kindly refer to the manual posted on the web. If your sample is with low concentration, there is indeed the possibility for an undetectable result. Usually, if the concentration is in the detection range of the kit, it can be detectable. Actually, the detection range of 62.5 pg/mL-4000 pg/mL means a more stable result in this range.
If your sample is indeed lower than the detection limit of the kit, very sorry there are no better solutions available so far.