Delve into the fascinating world of DNA repair and recombination research with the Recombinant Saccharomyces cerevisiae RAD52. This product, derived from the well-established model organism Baker's yeast (Saccharomyces cerevisiae, strain ATCC 204508 / S288c), presents a remarkable tool for studying DNA damage response and its role in genetic stability. The protein encoded by the RAD52 gene plays a crucial part in homologous recombination repair, making it a subject of keen interest in genetics and molecular biology.
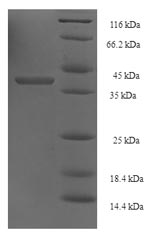
This recombinant RAD52 variant encompasses a partial sequence (60-282 amino acids), conserving essential functionalities to facilitate your research endeavors. The protein, expressed in E.coli, carries an N-terminal 6xHis-SUMO tag to facilitate its easy identification and purification. The SDS-PAGE analysis vouches for its high purity level, exceeding 90%. Choose between liquid or lyophilized powder form based on your lab's requirements. With the Recombinant Saccharomyces cerevisiae RAD52, you're not just purchasing a product; you're ensuring precise, high-quality, and reliable results for your research.