The SARS-CoV-2 spike glycoprotein receptor-binding domain represents a critical interface in viral pathogenesis, mediating the initial attachment to human ACE2 receptors that enables cellular entry. This interaction has made the RBD a focal point for therapeutic development, diagnostic assay design, and fundamental studies of coronavirus biology.
This recombinant nanobody, derived from alpaca VHH sequences fused to a human IgG1 Fc domain, offers researchers a uniquely compact binding format with exceptional affinity. The recombinant production method ensures sequence-defined consistency between lots, eliminating the batch variability that can complicate longitudinal studies or assay standardization. Affinity chromatography purification delivers a reagent ready for demanding applications.
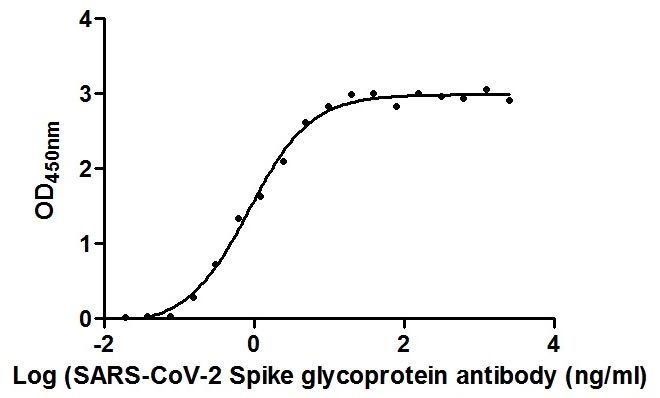
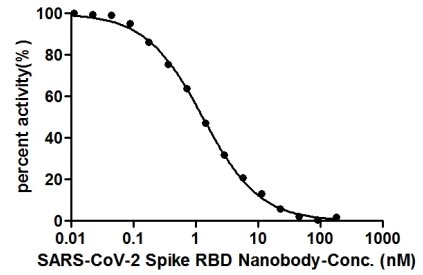
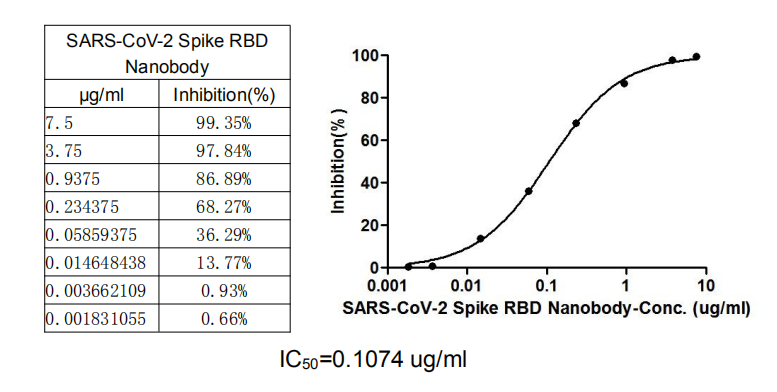
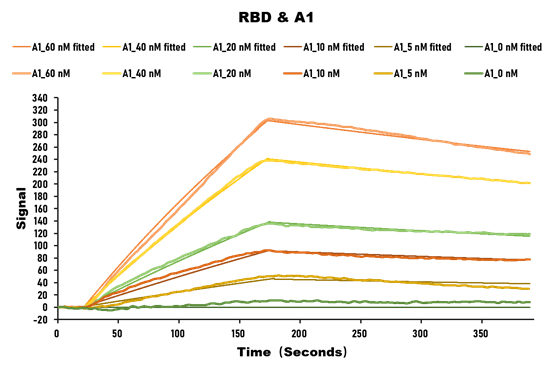
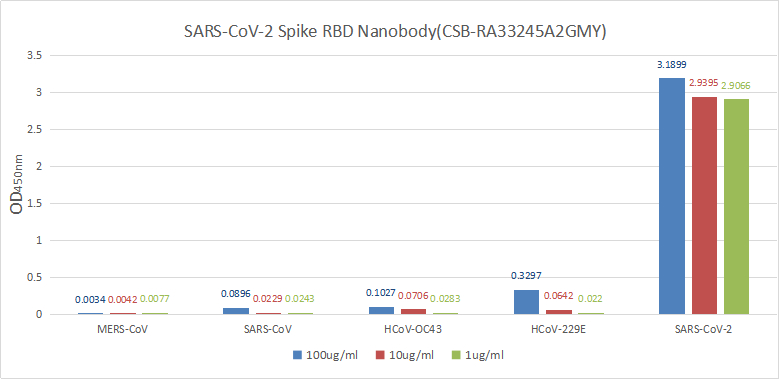
Validation data demonstrate impressive binding characteristics, with an EC50 of 0.8674 ng/ml in functional ELISA against immobilized S1-RBD and a measured affinity constant of 28.2 nM by surface plasmon resonance. Critically for neutralization studies, this nanobody effectively blocks the RBD-ACE2 interaction with an IC50 of 1.296 nM, providing a valuable tool for investigating viral entry mechanisms or developing competitive binding assays. Specificity testing confirms reactivity exclusively with SARS-CoV-2 S1-RBD, with no detectable binding to spike proteins from MERS-CoV, SARS-CoV, HCoV-OC43, or HCoV-229E, ensuring confidence in coronavirus-specific applications.
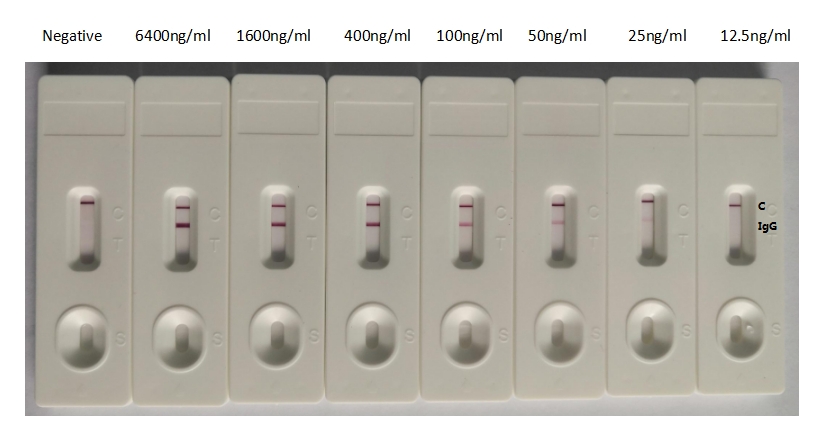
The antibody performs across multiple platforms including ELISA at dilutions from 1:10,000 to 1:100,000, colloidal gold immunochromatography assays with detection limits as low as 25 ng/ml, and neutralization studies. This versatility supports workflows ranging from high-throughput screening to rapid diagnostic development, making it a practical choice for virology research, vaccine evaluation studies, and infectious disease diagnostics.