With the COVID-19 spreads around the world, scientists have work hard to explore the mechanism of SARS-CoV-2 invasion. And there are many SARS-CoV-2-Host interactome targets which have been reported, such as ACE2, TMPRSS2 and CD147. The relationship between SARS-CoV-2 and TMPRSS2 was firstly reported in the study entitled “The novel coronavirus 2019 (2019-nCoV) uses the SARS-coronavirus receptor 1 ACE2 and the cellular protease TMPRSS2 for entry into target cells” [1]. Then, more and more studies about TMPRSS2 have been published. So what is the TMPRSS2? And how does SARS-CoV-2 invade human body via TMPRSS2?
1. What is The TMPRSS2?
Transmembrane protease serine 2, also known as TMPRSS2, is a member of transmembrane protease serines (TMPRSSs), which are a family of proteins with conserved serine protease domains located on the cell membrane. The basic structure of TMPRSSs is similar. The C-terminal protease domain is outside the cell, the N-terminal is located inside the cell, and it also has a single transmembrane domain. The difference lies in the backbone region.
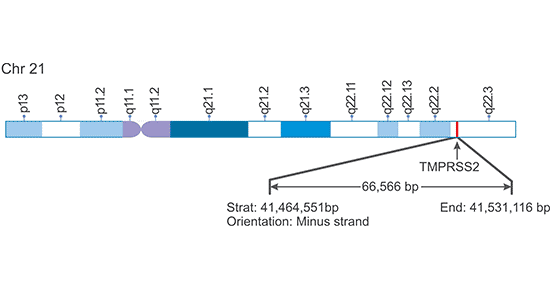
As the Figure 1 shows, TMPRSS2 gene is located on human chromosome 21: 41, 464, 551-41, 531, 116. A significant feature of the TMPRSS2 gene is that several androgen receptor elements (AREs) are located upstream of the transcription start site and the first intron [2] [3].
Figure 1. A schematic diagram of TMPRSS2 genomic location
*This diagram is derived from the publication published on Biochimie [2]
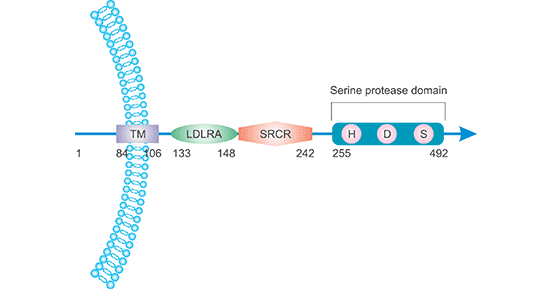
The TMPRSS2 protein, encoded by TMPRSS2 gene, consists of 492 amino acids which anchors to the plasma membrane. As the Figure 2 shows, it converts to its form through autocatalytic cleavage between Arg255 and Ile256. After cleavage, the mature proteases are mostly membrane-bound, yet a noticeable portion of them can be liberated into the extracellular milieu. The protease catalytic domain contains a catalytic triad consisting of the amino acid residues His296, Asp345 and Ser441, corresponding to His57, Asp102 and Ser195 of chymotrypsinogen [4].
Figure 2. The location and structure of TMPRSS2 protein
*This diagram is derived from the publication published on Biochimie [2]
2. What is The Distribution of TMPRSS2?
The expression of TMPRSS2 has obvious tissue specificity. Human TMPRSS2 is an androgen-regulated, type II transmembrane-bound serine protease that is predominantly expressed in prostate, with relatively lower level of expression in lungs, colon, liver, kidneys and pancreas. According to in situ hybridization analysis of mouse embryos and adult tissues, mouse TMPRSS2 is also expressed in gastrointestinal tract, urogenital tract and respiratory tract epithelial cells, indicating that the expression distribution of TMPRSS2 in mice and humans is very similar. The tissue specific expression also suggests that diseases caused by abnormal TMPRSS2 may prefer to men than women.
3. What is The Function of TMPRSS2?
TMPRSS2 is closely related to prostate cancer. In 2005, most prostate cancers (up to 70%) were found to have fusion of the TMPRSS2 and oestrogen-regulated gene (ERG), both on chromosome 21. Soon thereafter, other members of erythroblast transformation-specific (ETS) variant gene (ETV) family were found to have gene fusions, although at much lower frequencies, including ETV1 (chromosome 7), ETV4 (chromosome 17), ETV5 (chromosome 3), and ETS domain-containing protein gene (ELK4, chromosome 1).
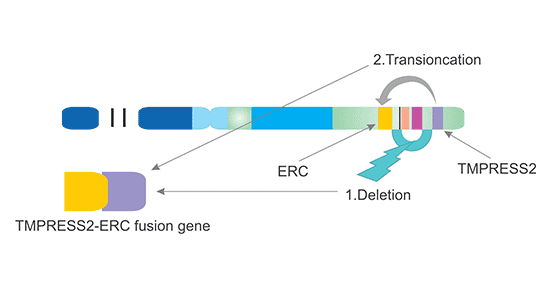
Taking the TMPRSS2 and ERG fusion as an example, gene fusion, also called gene rearrangement, often causes abnormal activation of certain genes. As the Figure 3 shows, the gene rearrangement consists of actual loss of genetic material between two genes on the same chromosome; note the deletion of intervening genes between TMPRSS2 and ERG on chromosome 21. A second form of fusion is due to translocation, when a gene moves to another location on the same chromosome or a different chromosome. Both mechanisms apply for the gene rearrangements in prostate cancer. Given the high prevalence of prostate cancer, this fusion gene is probably the most common fusion gene in human cancer. Further, TMPRSS2-ERG can be used as a diagnostic marker for prostate cancer.
Figure 2. Mechanism of TMPRESS2-ERG fusion (chromosome 21)
*This diagram is derived from the publication published on BJU Int. [5]
Additionally, besides its role in prostate cancer, it also related to virus. We will illustrate this function in the next section.
4. How is TMPRSS2 Related to Viruses?
In 2019, the study from Naoko Iwata-Yoshikawa et al. showed that TMPRSS2 activates the spike protein of highly pathogenic human coronavirus, such as severe acute respiratory syndrome-related coronavirus (SARS-CoV) and Middle East respiratory syndrome-related coronavirus (MERS-CoV). In vitro, the activation of TMPRSS2 induces virus-cell membrane fusion at the cell surface. In this study, they examined the role of TMPRSS2 using animal models of SARS-CoV and MERS-CoV infection. The results showed that lack of TMPRSS2 in the airways reduces the severity of lung pathology after infection by SARS-CoV and MERS-CoV [6].
Actually, as early as 2011, Ilona Glowacka and others from the Hannover Medical School in German assessed whether the S (Spike glycoprotein) of SARS-CoV is proteolytically processed by TMPRSS2. In this study, the results of western blot analysis revealed that SARS S was cleaved into several fragments upon coexpression of TMPRSS2 (cis-cleavage) and upon contact between SARS S-expressing cells and TMPRSS2-positive cells (trans-cleavage). cis-cleavage resulted in release of SARS S fragments into the cellular supernatant and in inhibiting antibody-mediated neutralization. trans-cleavage activated SARS S on effector cells to fuse with target cells. It shows that TMPRSS2 may promote viral spread and pathogenesis by reducing viral recognition by neutralizing antibodies and by activating SARS S for cell-cell and virus-cell fusion [7].
5. How does SARS-CoV-2 Invade Human Body via TMPRSS2?
SARS-CoV-2, one type of coronavirus, share the central biological properties and similar structure with SARS-CoV. Regarding the ways of coronavirus to enter cells involving TMPRSS2, two studies reported on two ways back in 2013. One is that the cell surface is mediated by TMPRSS2, and the other is mediated by cathepsin L in the nucleus. Stefanie Gierer et al. found that TMPRSS2 and cathepsins B and L can activate the novel human coronavirus EMC (hCoV-EMC) and fuse with target cells. Therefore, TMPRSS2 and cathepsin have become potential targets for controlling hCoV-EMC [8] [9]. In 2014, Adeline Heurich and others once again proved the role of TMPRSS2 and HAT in coronavirus. They found that these two enzymes can cleave and activate the spike protein of SARS-CoV for membrane fusion. In addition, these enzymes also cleave SARS-CoV receptor ACE2 [10].
In 2020, with the COVID-19 globally broke out, Markus Hoffmann and other further to demonstrate that coronaviruses use their spike proteins to select and enter target cells and insights into SARS-CoV-2 spike (S)-driven entry might facilitate assessment of pandemic potential and reveal therapeutic targets. In their study, they demonstrated that SARS-CoV-2-S uses the SARS-CoV receptor, ACE2, for entry and the cellular protease TMPRSS2 for SARS-CoV-2-S priming. A TMPRSS2 inhibitor blocked entry and might constitute a treatment option. These results revealed important commonalities between SARS-CoV-2 and SARS-CoV infection, which might translate into similar transmissibility and disease pathogenesis [1] [11].
6. The Latest Progression of TMPRSS2 Research
In this section, we list several latest progression of TMPRSS2 and SARS-CoV-2 Research as follows:
- On April 12, 2021, the team from Stanford University and University of Iowa created a structure-based phylogenetic computational tool named 3DPhyloFold to systematically identify structurally similar serine proteases (TMPRSS2) with known therapeutic inhibitors and demonstrated effective inhibition of SARS-CoV-2 infection in vitro and in vivo [12].
- On Mar 19, 2021, the team from Ulm University Medical Center reported that the acute phase protein α1AT is an inhibitor of TMPRSS2 and SARS-CoV-2 entry, and may play an important role in the innate immune defense against the novel coronavirus [13].
- On Mar 11, 2021, the team from Chang Gung University revealed that risk of SARS-CoV-2 infection and COVID-19 disease severity increased by air pollution exposure and underlying IPF. It can be mediated through upregulating ACE2 and TMPRSS2 in pulmonary fibroblasts, and prevented by blocking the IL-8/CXCR1/2 pathway [14].
- On Jan 26, 2021, the team from Fondazione IRCCS Policlinico San Matteo demonstrated that MSCs derived from different human tissues are not permissive to SARS-CoV-2 infection, support the safety of MSCs as potential therapy for COVID-19 [15].
- In Jan, 2021, the research data of the team from Hokkaido University were shown to result from their inability to utilize the entry pathway involving direct fusion mediated by the host type II transmembrane serine protease, TMPRSS2. The study demonstrated that the S protein polybasic cleavage motif is a critical factor underlying SARS-CoV-2 entry and cell tropism [16].
7. The drugs Targeting TMPRSS2 of SARS-CoV-2 Treatment
Accumulating evidence has shown that TMPRSS2 is a powerful target in SARS-CoV-2 (COVID 19) treatment. We collect the latest clinic data of drugs targeting TMPRSS2 of SARS-CoV-2 treatment as follows:
| Title |
Types of drugs |
Indications |
Phase |
Sponsors |
Last update date |
| Low-dose Hydroxychloroquine and Bromhexine: a Novel Regimen for COVID-19 Prophylaxis in Healthcare Professionals (ELEVATE Trial) |
Organic heterocyclic drugs |
Hydroxychloroquine; Antimalarials; Enzyme Inhibitors; Antirheumatic Agents |
Early Phase 1 |
Instituto Nacional de Rehabilitacion |
April 5, 2021 |
| Randomized Trial of Bicalutamide to Block TMPRSS2 in Males With COVID-19 Infection |
Organic heterocyclic drugs |
COVID-19 |
Phase 3 |
University of Florida |
January 27, 2021 |
| RECOVER: Phase 2 Randomized, Double-Blind Trial TREating Hospitalized Patients With COVID-19 With Camostat MesilatE, a TMPRSS2 Inhibitor |
Organic heterocyclic drugs |
Severe Acute Respiratory Syndrome |
Phase 2 |
Alan Bryce |
January 11, 2021 |
| An Exploratory Analysis of the Expression of Receptors and Activating Proteases Mediating SARS-CoV-2 Entry and the Association Between HSD3B1 Gene Polymorphisms With Outcomes in SARS-CoV-2 Infected Patients |
/ |
Covid19 |
N/A |
Ricardo Pereira Mestre |
August 18, 2020 |
| RAndomized Clinical Trial in COvid19 Patients to Assess the Efficacy of the Transmembrane Protease Serine 2 (TMPRSS2) Inhibitor NAfamostat (RACONA Study) |
Organic heterocyclic drugs |
COVID19 |
Phase 2/Phase 3 |
University Hospital Padova |
April 20, 2020 |
| Efficacy of Aerosol Combination Therapy of 13 Cis Retinoic Acid and Captopril for Treating Covid-19 Patients Via Indirect Inhibition of Transmembrane Protease, Serine 2 (TMPRSS2) |
Organic heterocyclic drugs |
Covid19 |
Phase 2 |
Kafrelsheikh University |
October 26, 2020 |
References
[1] Markus Hoffmann, Hannah Kleine-Weber, et al. The novel coronavirus 2019 (2019-nCoV) uses the SARS-coronavirus receptor ACE2 and the cellular protease TMPRSS2 for entry into target cells [J]. bioRxiv. 2020.
[2] Li Wen Shen, Hui Juan Mao, et al. TMPRSS2: A potential target for treatment of influenza virus and coronavirus infections [J]. Biochimie. 2017.
[3] Y. Park. TMPRSS2 (transmembrane protease, serine 2) [J]. Atlas. Genet. Cytogenet. Oncol. Haematol. 2010, 14: 1163−1165.
[4] B. Lin, C. Ferguson, et al. Prostate-localized and androgen-regulated expression of the membrane-bound serine protease TMPRSS2 [J]. Cancer Res. 1999, 59:4180−4184.
[5] Deloar Hossain and David G. Bostwick. Significance of the TMPRSS2: ERG gene fusion in prostate cancer [J]. BJU Int. 2013.
[6] Naoko Iwata-Yoshikawa, Tadashi Okamura, et al. TMPRSS2 Contributes to Virus Spread and Immunopathology in the Airways of Murine Models After Coronavirus Infection [J]. J Virol. 2019, 93(6):e01815-18.
[7] Ilona Glowacka, Stephanie Bertram, et al. Evidence That TMPRSS2 Activates the Severe Acute Respiratory Syndrome Coronavirus Spike Protein for Membrane Fusion and Reduces Viral Control by the Humoral Immune Response [J]. J Virol. 2011, 85(9):4122-34.
[8] Gierer S, Bertram S, et al. The spike protein of the emerging betacoronavirus EMC uses a novel coronavirus receptor for entry, can be activated by TMPRSS2, and is targeted by neutralizing antibodies [J]. J Virol. 2013, 87(10):5502-11.
[9] Bertram S, Dijkman R, et al. TMPRSS2 activates the human coronavirus 229E for cathepsin-independent host cell entry and is expressed in viral target cells in the respiratory epithelium [J]. J Virol. 2013, 87(11):6150-60.
[10] Heurich A, Hofmann-Winkler H, et al. TMPRSS2 and ADAM17 cleave ACE2 differentially and only proteolysis by TMPRSS2 augments entry driven by the severe acute respiratory syndrome coronavirus spike protein [J]. J Virol. 2014, 88(2):1293-307.
[11] Markus Hoffmann, Hannah Kleine-Weber, et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor [J]. Cell. 2020, 181(2):271-280.
[12] Sun YJ, Velez G, Parsons DE, et al. Structure-based phylogeny identifies Avoralstat as a TMPRSS2 inhibitor that prevents SARS-CoV-2 infection in mice [J]. J Clin Invest. 2021, 147973.
[13] Wettstein L, Weil T, Conzelmann C, et al. Alpha-1 antitrypsin inhibits TMPRSS2 protease activity and SARS-CoV-2 infection [J]. Nat Commun. 2021;12(1):1726.
[14] Li HH, Liu CC, Hsu TW, et al. Upregulation of ACE2 and TMPRSS2 by particulate matter and idiopathic pulmonary fibrosis: a potential role in severe COVID-19 [J]. Part Fibre Toxicol. 2021, 18(1):11.
[15] Avanzini MA, Mura M, Percivalle E, et al. Human mesenchymal stromal cells do not express ACE2 and TMPRSS2 and are not permissive to SARS-CoV-2 infection [J]. Stem Cells Transl Med. 2021, 10(4):636-642.
[16] Sasaki M, Uemura K, Sato A, et al. SARS-CoV-2 variants with mutations at the S1/S2 cleavage site are generated in vitro during propagation in TMPRSS2-deficient cells [J]. PLoS Pathog. 2021, 17(1):e1009233.
CUSABIO team. TMPRSS2, an Activator for SARS-CoV-2 Invasion. https://www.cusabio.com/c-20986.html



Comments
Leave a Comment