The SARS-CoV-2 spike glycoprotein serves as the primary mediator of viral entry into host cells, making it a central focus for researchers studying coronavirus biology, vaccine development, and therapeutic interventions. The receptor-binding domain within the S1 subunit directly engages human ACE2, establishing this protein as a critical target for neutralization studies and diagnostic assay development.
This recombinant monoclonal antibody, clone H6, offers researchers the reproducibility essential for longitudinal studies and multi-site collaborations. As a sequence-defined reagent produced through recombinant technology, it eliminates the lot-to-lot variability inherent in traditional hybridoma-derived antibodies, ensuring your experimental conditions remain consistent across projects.
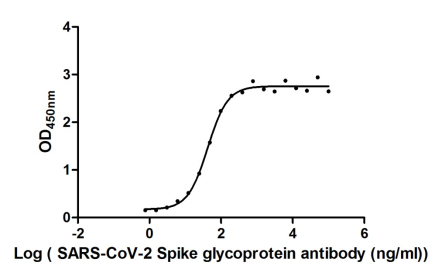
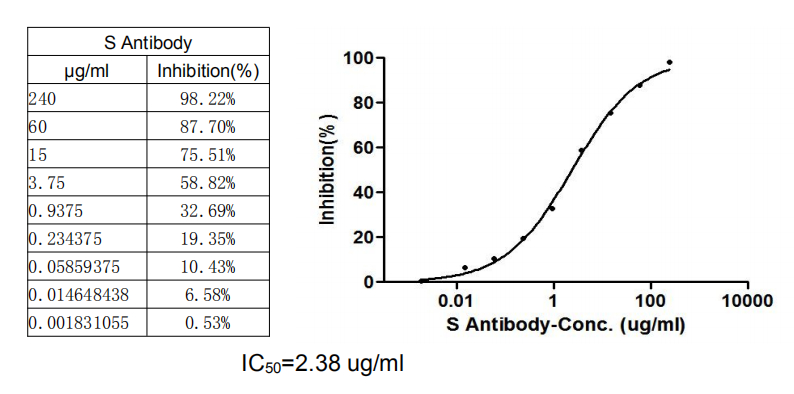
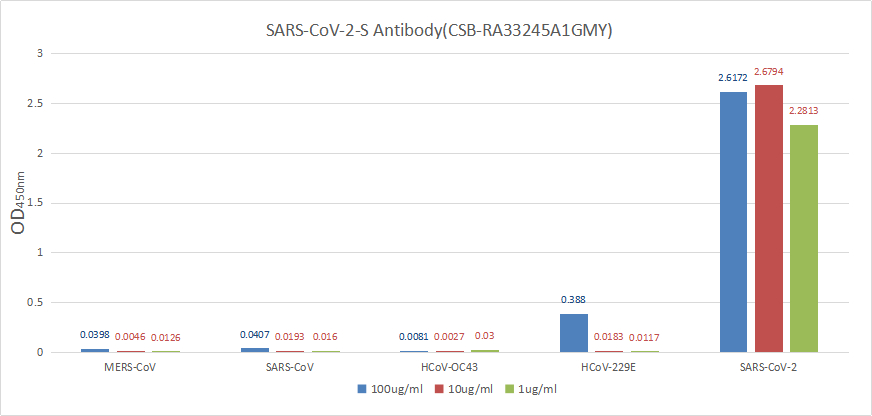
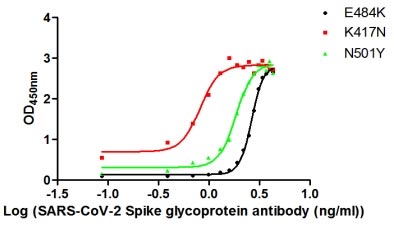
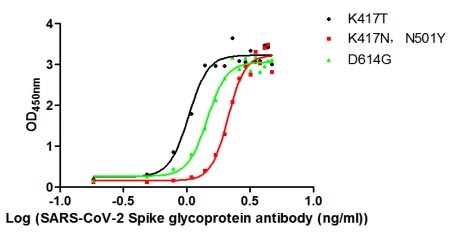
Functional validation demonstrates robust binding characteristics, with EC50 values of 42.83 ng/ml against the full spike protein and 29.51 ng/ml against the S1-RBD specifically. Importantly, this antibody exhibits genuine neutralizing capability, blocking the spike-ACE2 interaction with an IC50 of 23.32 nM, making it suitable for functional studies examining viral entry mechanisms. Specificity testing confirms reactivity is restricted to SARS-CoV-2, with no cross-reactivity observed against MERS-CoV, SARS-CoV, HCoV-OC43, or HCoV-229E spike proteins—a valuable characteristic when distinguishing between coronavirus strains in your research.
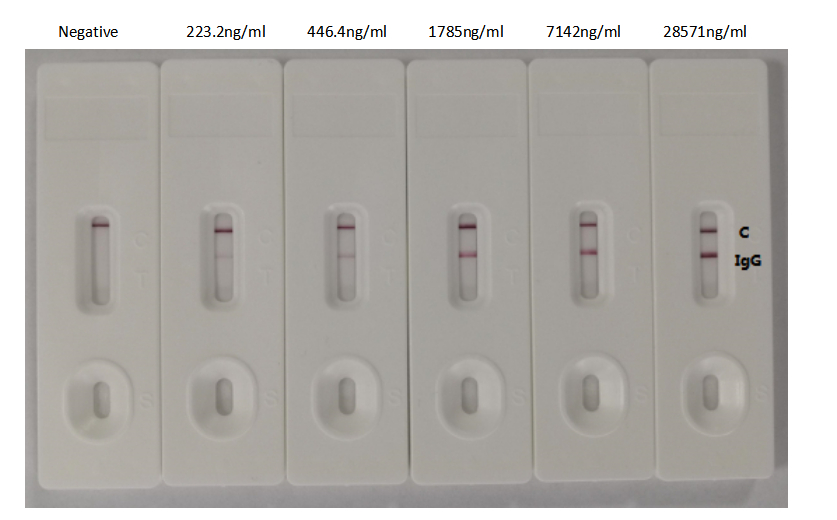
The antibody performs across multiple platforms, including ELISA at dilutions from 1:10,000 to 1:50,000, colloidal gold immunochromatography assays with detection limits as low as 223.2 ng/ml, and neutralization assays. This versatility supports workflows ranging from quantitative binding studies to rapid diagnostic development. The scFv-human IgG1 Fc fusion format combines the specificity of a defined binding domain with the stability and detection compatibility of a human constant region, supporting diverse downstream applications in infectious disease research.