The Human 8-oxoguanine DNA glycosydase (hOGG1) ELISA kit is designed to enable accurate and reliable detection of 8-oxoguanine DNA glycosydase in Homo sapiens (Human) samples.
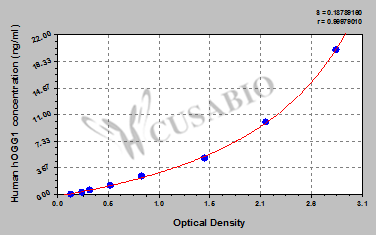
This ELISA kit offers a versatile solution for researchers studying cancer、DNA repair, allowing for the detection of hOGG1 in serum, tissue homogenates, and cell lysates. With a wide detection range of 0.312 ng/mL-20 ng/mL and a sensitivity of 0.078 ng/mL, this kit provides accurate and sensitive measurements of hOGG1 levels in samples.
The assay time for this ELISA kit is 1-5 hours, and sample volume ranges from 50-100ul. The detection wavelength is 450 nm, and the measurement method is sandwich, providing a quantitative approach for researchers studying the role of hOGG1 in cancer and DNA repair