Function
Mono-ADP-ribosyltransferase that mediates mono-ADP-ribosylation of target proteins and plays a key role in the response to DNA damage. Mediates mono-ADP-ribosylation of glutamate, aspartate or lysine residues on target proteins. In contrast to PARP1 and PARP2, it is not able to mediate poly-ADP-ribosylation. Associates with a number of DNA repair factors and is involved in the response to exogenous and endogenous DNA strand breaks. Together with APLF, promotes the retention of the LIG4-XRCC4 complex on chromatin and accelerate DNA ligation during non-homologous end-joining (NHEJ). Cooperates with the XRCC5-XRCC6 (Ku80-Ku70) heterodimer to limit end-resection thereby promoting accurate NHEJ. Involved in DNA repair by mediating mono-ADP-ribosylation of a limited number of acceptor proteins involved in chromatin architecture and in DNA metabolism, such as XRCC5 and XRCC6. ADP-ribosylation follows DNA damage and appears as an obligatory step in a detection/signaling pathway leading to the reparation of DNA strand breaks. May link the DNA damage surveillance network to the mitotic fidelity checkpoint. In addition to proteins, also able to ADP-ribosylate DNA: mediates DNA mono-ADP-ribosylation of DNA strand break termini via covalent addition of a single ADP-ribose moiety to a 5'- or 3'-terminal phosphate residues in DNA containing multiple strand breaks. Acts as a negative regulator of immunoglobulin class switch recombination, probably by controlling the level of AICDA /AID on the chromatin.
Subcellular Location
Nucleus. Chromosome. Cytoplasm, cytoskeleton, microtubule organizing center, centrosome. Cytoplasm, cytoskeleton, microtubule organizing center, centrosome, centriole.
Tissue Specificity
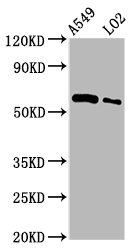
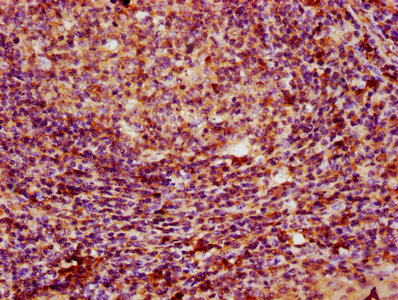
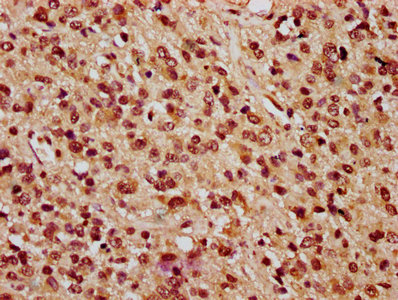
Widely expressed; the highest levels are in the kidney, skeletal muscle, liver, heart and spleen; also detected in pancreas, lung, placenta, brain, leukocytes, colon, small intestine, ovary, testis, prostate and thymus.