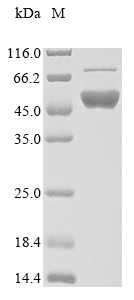
The region for expressing recombinant Human FEN1 contains amino acids 1-380. This FEN1 protein is expected to have a theoretical molecular weight of 50.0 kDa. Expression of this FEN1 protein is conducted in e.coli. The FEN1 gene fragment has been modified by fusing the N-terminal 10xHis tag and C-terminal Myc tag, providing convenience in detecting and purifying the recombinant FEN1 protein during the following stages.
Flap endonuclease 1 (FEN1) is a multifunctional endonuclease involved in DNA replication and repair processes. In humans, FEN1 plays a crucial role in processing DNA intermediates during Okazaki fragment maturation, long-patch base excision repair, and nucleotide excision repair. As a structure-specific nuclease, FEN1 cleaves 5' flaps, flap structures that arise during DNA synthesis. This activity is vital for maintaining genomic stability and integrity. FEN1 also participates in DNA damage response pathways, contributing to the repair of various lesions. Dysregulation of FEN1 has been associated with genomic instability and is implicated in cancer development. Understanding FEN1's functions is essential for unraveling DNA repair mechanisms and potential therapeutic targets for cancer treatment.