SMC1A plays a central role in the cohesin complex, which is essential for proper chromosome segregation during cell division and DNA damage repair. Phosphorylation at serine 957 represents a critical regulatory modification that occurs in response to DNA damage, particularly through the ATM kinase pathway. Detecting this specific phosphorylation event allows researchers to monitor DNA damage checkpoint activation and study the intricate signaling networks that maintain genomic stability.
This recombinant monoclonal antibody, clone 1F9, offers the reproducibility and consistency that phospho-specific detection demands. Because the antibody sequence is defined and produced recombinantly in rabbit, researchers can expect uniform performance across experiments and over time, eliminating the lot-to-lot variability that can complicate longitudinal studies or multi-site collaborations. The affinity-chromatography purification ensures high specificity for the phosphorylated epitope.
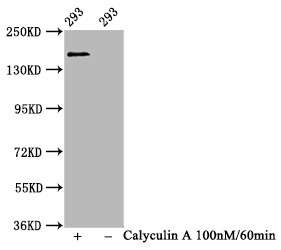
Validation by western blot demonstrates reliable detection in human 293 whole cell lysates, with the antibody successfully distinguishing between untreated cells and those treated with Calyculin A, a phosphatase inhibitor that enhances phosphorylation signals. The observed band at 160 kDa matches the predicted molecular weight precisely, confirming target specificity. For western blot applications, dilutions ranging from 1:500 to 1:5000 provide flexibility to optimize signal intensity based on your experimental conditions and sample phosphorylation levels. The antibody is also validated for ELISA, expanding its utility across different experimental workflows.
For researchers investigating DNA damage responses, cell cycle checkpoints, or cohesin biology, this phospho-SMC1A antibody provides a dependable tool for tracking this functionally significant post-translational modification in human samples.