The SARS-CoV-2 Spike glycoprotein remains a critical focus in infectious disease research, serving as the primary mediator of viral entry through its interaction with the human ACE2 receptor. The receptor-binding domain within the S1 subunit represents a key target for understanding viral pathogenesis, evaluating neutralizing antibody responses, and developing diagnostic assays. This biotin-conjugated recombinant monoclonal antibody provides researchers with a reliable tool for detecting and quantifying Spike protein in ELISA-based workflows.
Developed using recombinant technology, this antibody offers the consistency that demanding research requires. The sequence-defined nature of recombinant production eliminates the lot-to-lot variability inherent in traditional hybridoma methods, ensuring that your experimental conditions remain reproducible across studies. The mouse scFv fusion with human IgG1 Fc architecture combines the specificity of the original clone with enhanced stability, while the biotin conjugation enables seamless integration into streptavidin-based detection systems without additional labeling steps.
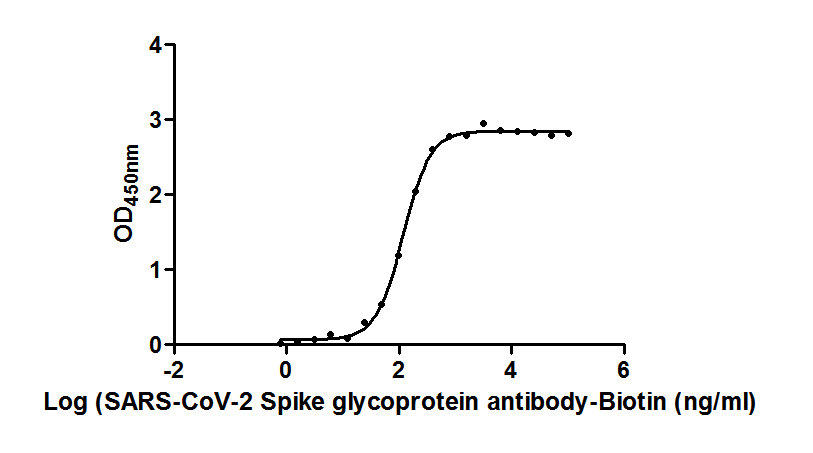
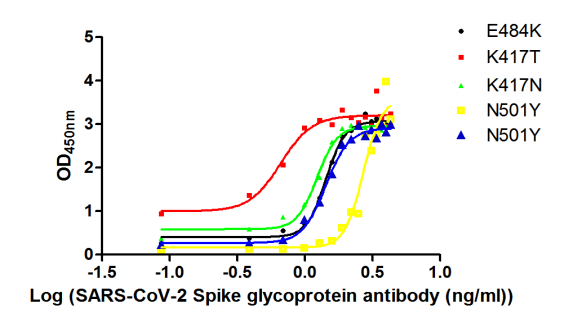
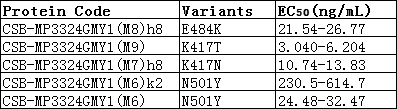
Functional ELISA validation demonstrates robust binding activity, with the antibody recognizing immobilized SARS-CoV-2 S1-RBD at an EC50 of 118.7 ng/ml. This sensitivity supports detection across a broad working range, with recommended dilutions spanning 1:10,000 to 1:50,000 depending on your specific assay conditions and antigen concentrations. The antibody was raised against a recombinant Spike glycoprotein fragment encompassing amino acids 16-685, covering the functionally critical S1 region.
For researchers investigating SARS-CoV-2 biology, vaccine development, or serological assay optimization, this biotin-conjugated antibody delivers the performance and reproducibility needed for meaningful results in coronavirus research applications.