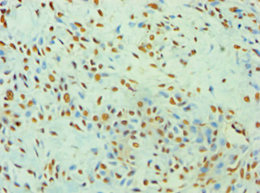
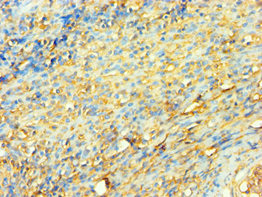
Single-stranded DNA-dependent ATP-dependent helicase that plays a key role in DNA non-homologous end joining (NHEJ) by recruiting DNA-PK to DNA. Required for double-strand break repair and V(D)J recombination. Also has a role in chromosome translocation. The DNA helicase II complex binds preferentially to fork-like ends of double-stranded DNA in a cell cycle-dependent manner. It works in the 3'-5' direction. During NHEJ, the XRCC5-XRRC6 dimer performs the recognition step: it recognizes and binds to the broken ends of the DNA and protects them from further resection. Binding to DNA may be mediated by XRCC6. The XRCC5-XRRC6 dimer acts as regulatory subunit of the DNA-dependent protein kinase complex DNA-PK by increasing the affinity of the catalytic subunit PRKDC to DNA by 100-fold. The XRCC5-XRRC6 dimer is probably involved in stabilizing broken DNA ends and bringing them together. The assembly of the DNA-PK complex to DNA ends is required for the NHEJ ligation step. The XRCC5-XRRC6 dimer probably also acts as a 5'-deoxyribose-5-phosphate lyase (5'-dRP lyase), by catalyzing the beta-elimination of the 5' deoxyribose-5-phosphate at an abasic site near double-strand breaks. XRCC5 probably acts as the catalytic subunit of 5'-dRP activity, and allows to 'clean' the termini of abasic sites, a class of nucleotide damage commonly associated with strand breaks, before such broken ends can be joined. The XRCC5-XRRC6 dimer together with APEX1 acts as a negative regulator of transcription. In association with NAA15, the XRCC5-XRRC6 dimer binds to the osteocalcin promoter and activates osteocalcin expression. As part of the DNA-PK complex, involved in the early steps of ribosome assembly by promoting the processing of precursor rRNA into mature 18S rRNA in the small-subunit processome. Binding to U3 small nucleolar RNA, recruits PRKDC and XRCC5/Ku86 to the small-subunit processome. Plays a role in the regulation of DNA virus-mediated innate immune response by assembling into the HDP-RNP complex, a complex that serves as a platform for IRF3 phosphorylation and subsequent innate immune response activation through the cGAS-STING pathway.