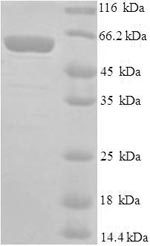
The process of expressing the recombinant mouse Mdm2 protein in the E.coli requires the recombinant DNA gene formed by the integration of encoding gene for the 1-489aa of the mouse Mdm2 protein and N-terminal 6xHis tag sequence, the expression vector that the recombinant DNA gene inserts into, the E.coli that provided the necessary macromolecules and components for transcription and translation of the cloned expression vector. After isolation and purification, this N-terminal 6xHis-tagged recombinant Mdm2 protein was obtained. This recombinant Mdm2 protein is characterized by high purity (>90%, SDS-PAGE). This Mdm2 protein ran along the gel to the band of approximately 58 kDa molecular weight.
E3 ubiquitin-protein ligase Mdm2 is a protein encoding by a gene named Mdm2 in mouse and a gene named MDM2 in human. Generally, MDM2 is known as a negative regulator of the tumour suppressor p53, making it an attractive target for anti-cancer drug design. Mdm2 affects p53 stability from two sections, one is as an E3 ubiquitin ligase that recognizes the N-terminal trans-activation domain of the p53 tumor suppressor, another is as an inhibitor of p53 transcriptional activation. Currently, diseases involved MDM2 include Lessel-Kubisch Syndrome and Accelerated Tumor Formation.