As mentioned in the article entitled "What a Powerful Artifact of DNA Compression - Histone", a histone modification is a series of post-translational modification (PTM) to histone proteins including methylation, phosphorylation, acetylation, ubiquitylation, and sumoylation. The PTMs made to histones can impact gene expression though altering chromatin structure or recruiting histone modifiers. But its mechanism is remain unclear.
In the nuclei of eukaryotic cells, DNA wraps around the octamer of histone proteins to form the nucleosome, in a structure like ‘beads on a string’, which makes up the basic unit of chromatin. Chromatin further folds into higher-level structures, loosely or tightly, which helps to determine the accessibility of the DNA.
Last year, two scientists were honored The 2018 Albert Lasker Basic Medical Research Award for discoveries, which have elucidated how gene expression is influenced by chemical modification of histones, the proteins that package DNA within chromosomes.
Michael Grunstein, a professor from University of California in Los Angeles, demonstrated that histones dramatically influence gene activity within living cells and laid the groundwork for understanding the pivotal role of particular amino acids in this process through tour-de-force genetic studies in yeast [1]. C. David Allis, a professor from Rockefeller University, uncovered an enzyme that attaches a specific chemical group to a particular amino acid in histones, and this histone-modifying enzyme turned out to be an established gene co-activator whose biochemical capabilities had eluded researchers [2]. Grunstein and Allis unveiled a previously hidden layer of gene control and broke open a new field.
Here, we have an example, histone acetylation. As it is shown in Figure 1. Transcriptional activation by histone acetylation, acetylated lysines which carry a positive charge, allow the histones to tightly bind DNA, which carries a negative charge. Consequently, the transcriptional machinery cannot access the DNA, and genes remain inactive. Addition of acetyl groups to particular lysines in histone tails neutralizes the positive charge and loosens the nucleosome's grip on DNA. This process allows the transcriptional machinery to access the DNA, and genes are active. Other modifications affect transcription in different ways [3] [4].
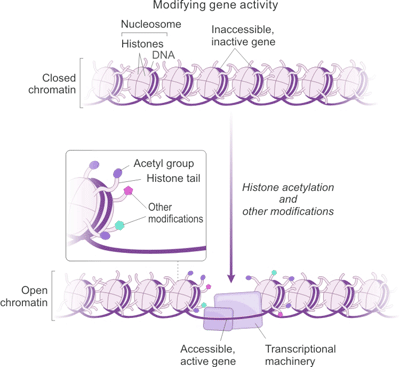
Figure 1. The Correlation between Histone Acetylation and Gene Expression
In summary, for the correlation between histone modification and gene expression, there is a word which can describe this kind of relationship very well, "transcriptional regulation is a complex and dynamic process of which histone modifications are a key component". Some histone modifications are established as consequences of active transcription; however, these histone modifications provide memory of recent transcriptional activity and signal for additional transcriptional regulation. Many histone modifications that are not coupled with active transcription function in transcriptional regulation processes. Rapid progress in genome-wide experiments, bioinformatics analysis and new technologies that allow epigenetic enzymes to be targeted to specified DNA sequences will help elucidate the genesis and biological functions of the myriad of histone modifications [5].
Click here to view the catalogue of Modified Histone Antibodies
References
[1] Shahbazian, M.D., and Grunstein, M. Functions of site-specific histone acetylation and deacetylation [J]. Annu. Rev. Biochem. 2007, 76, 75-100.
[2] Lewis, P.W., Mueller, M.M., et al. Inhibition of PRC2 activity by gain-of-function mutations in pediatric gliobastoma [J]. Science. 2013, 340, 857-861.
[3] Kurdistani, S.K., Tavazoie, S, et al. Mapping global histone acetylation patterns to gene expression [J]. Cell. 2004, 117, 721-733.
[4] Maze, I., Noh, K.M., et al. Every amino acid matters: essential contributions of histone variants to mammalian development and disease [J]. Nat. Rev. Genet. 2014, 15, 259-271.
[5] Xianjun Dong and Zhiping Weng. The correlation between histone modifications and gene expression [J]. Epigenomics. 2013 Apr; 5(2): 113–116.
CUSABIO team. How Does Histone Modification Affect Gene Expression?. https://www.cusabio.com/c-20826.html




Comments
Leave a Comment