Currently, in the research of histone modification, the most frequent and important issue is how to measure histone modification. And compared with the analysis of DNA methylomes, the implementation of epigenomic technologies for the study of histone modifications presents greater challenges than exist for the analysis of DNA methylomes. In this section, we introduce three main methods for detecting histone modification, including ChIP-on-chip, ChIP–SAGE (Chromatin immunoprecipitation combined with serial analysis of gene expression) and ChIP–Seq (Chromatin immunoprecipitation combined with high-throughput sequencing techniques).
Generally speaking, all of the three methods have a common method which is Chromatin immunoprecipitation assay (ChIP), then associated DNA can be investigated in several ways. For ChIP, there are two types ChIP, Native ChIP and Cross-linking ChIP, respectively. Native ChIP uses native chromatin prepared by nuclease digestion of cell nuclei. Cross-linking ChIP uses chromatin fixed with formaldehyde and fragmented by sonication. In the table 1, we have a comparison between the two ways of ChIP.
Table 1. The Comparison Between Native ChIP And Cross-linking ChIP
|
Types
|
advantages
|
disadvantages
|
|
Native ChIP
|
-
Antisera are generally raised against unfixed peptides or proteins, so the specificity of the antibody binding to unfixed chromatin is more predictable.
-
The immunoprecipitation is very efficient, usually yielding sufficient material for protein analysis.
|
-
NchIP is generally not applicable to non-histone proteins, although proteins that bind tightly to DNA can be studied.
-
Danger of protein rearrangement during chromatin preparation and precipitation;
-
There may be selective digestion of particular chromatin domains during preparation.
|
|
Cross-linking ChIP
|
-
Can be used to study non-histone proteins, even ones that do not, themselves, bind DNA.
-
Generally requires smaller numbers of cells than NChIP and less antibody.
-
Minimises the chances of chromatin rearrangements during preparation and precipitation.
|
-
Precipitation is often very inefficient, so insufficient protein is recovered to test the specificity of the precipitation reaction.
-
For at least some applications, the cross-linked protein--DNA complex must be purified by isopycnic centrifugation through cesium chloride, a prolonged and expensive procedure.
-
There is a danger that the cross-linking step may fix interactions that are transient and of minor functional significance.
|
Besides, you can click the following name of protocols to view the entire protocols.
Native Chromatin immunoprecipitation (ChIP) Protocol
Cross-linking Chromatin immunoprecipitation (ChIP) Protocol
1. ChIP-on-chip
ChIP-on-chip, also known as ChIP-chip, is a technology that combines chromatin immunoprecipitation with DNA microarray. Like regular ChIP, ChIP-on-chip is used to investigate interactions between proteins and DNA in vivo. Actually, the ChIP–chip method can be used to study many of the epigenomic phenomena. The example emerged here shows the application of ChIP–chip in studying histone modifications. As the Figure 1 shows:
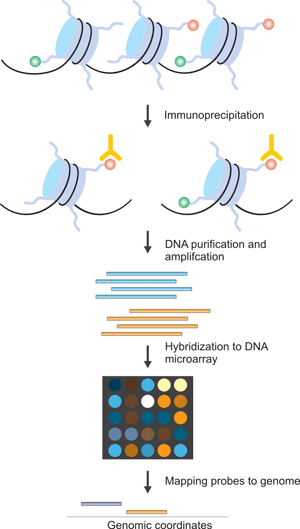
Figure 1. The Brief Processes of ChIP-on-chip
Modified chromatin is first purified by immunoprecipitating cross-linking chromatin with an antibody that is specific to a particular histone modification. Then DNA is amplified to obtain sufficient DNA (labelled with one color). The color-labelled ChIP DNA is hybridized to a DNA microarray with the control DNA prepared from input chromatin and labelled with a different color. Subsequently, the microarray probes can be mapped to the genome to yield genomic coordinates. However, most ChIP on chip experiments still use polyclonal antibodies, which are different in specificity between batches. The generation and use of monoclonal antibodies for ChIP on chip is therefore an important goal.
2. ChIP–SAGE
Additionally, combination of ChIP experiments with SAGE can profile histone modifications at a genomic scale. The ChIP–SAGE procedure begins with a ChIP step to purify chromatin regions that are associated with a specific histone modification, and proceeds as follows. It is shown in Figure 2:
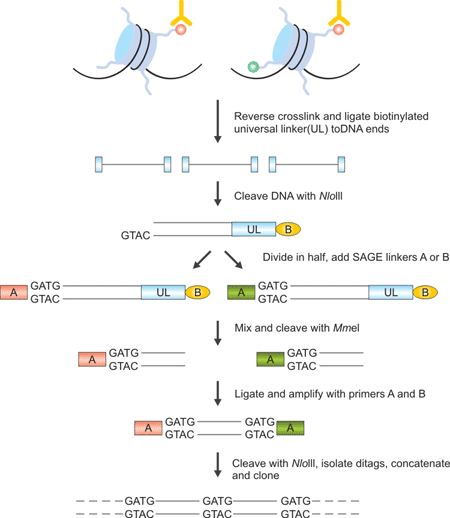
Figure 2. The Brief Processes of ChIP–SAGE
First of all, crosslinks are reversed, a biotinylated universal linker (UL) is ligated to DNA ends and DNA is bound to streptavidin beads. Then Nla III, which recognizes CATG, is used to digest DNA and a linker containing the recognition sequence of Mme I is ligated to the cleaved DNA ends. Mme I digestion produces 21–22 bp sequence tags from the immunoprecipitated fragments; the sequence tags are concatenated, cloned into a sequencing vector and sequenced. About 20 to 30 short sequence tags of 21 bp can be generated from each sequencing reaction. The sequence tags can then be mapped to the genome to identify modified regions.
3. ChIP–Seq
ChIP-seq, also known as ChIP-sequencing, is one of the most exciting recent advances in technologies for studying epigenetic phenomena at a genomic scale which relies on the combination of ChIP experiments with highthroughput sequencing. ChIP-seq combines ChIP with massively parallel DNA sequencing to identify the binding sites of DNA-associated proteins. As the Figure 3 shows:
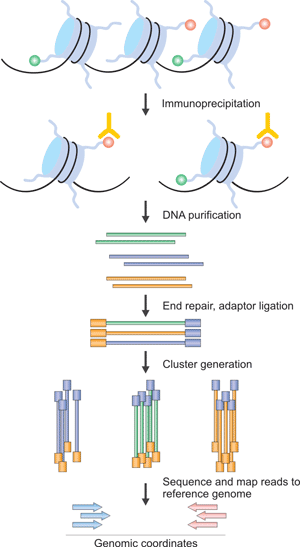
Figure 3. The Brief Processes of ChIP–Seq
The procedure that is outlined here is specific to the Illumina Genome Analyzer using Solexa technology. It can be divided into two part, ChIP and Sequencing. The first step is the purification of modified chromatin by immunoprecipitation with an antibody that is specific to a particular histone modification. The ChIP DNA ends are repaired and ligated to a pair of adaptors, followed by limited PCR amplification. The DNA molecules are bound to the surface of a flow cell that contains covalently bound oligonucleotides that recognize the adaptor sequences. After size selection, all the resulting ChIP-DNA fragments are sequenced simultaneously using a genome sequencer. The resulting sequence reads are mapped to a reference genome to obtain genomic coordinates that correspond to the immunoprecipitated fragments.
Click here to view the catalogue of Modified Histone Antibodies
CUSABIO team. How to Measure Histone Modification?. https://www.cusabio.com/c-20827.html






Comments
Leave a Comment