Currently, malignant tumor has become one of the causes of death. The incidence of cancer is increasing globally and the age of onset is getting younger.
Targeted therapies selectively kill cancer cells without affecting healthy tissue. Targeted drug development will be an important means to treat cancer.
The rapid development of high-throughput assay technology has led to the rapidaccumulation of tumor-related omics data. These data are of great significance for the study of the mechanism of tumor occurrence and development.
How to effectively use and store this information is especially important. The establishment of tumor bioinformatics database provides an effective solution, which has greatly promoted the development of basic tumor research and the improvement of clinical treatment level.
The following are some tumor-related database classifications and overviews:
References
[1] Samur M K, Yan Z, Wang X, et al. canEvolve: A Web Portal for Integrative Oncogenomics [J]. PLOS ONE, 2013, 8.
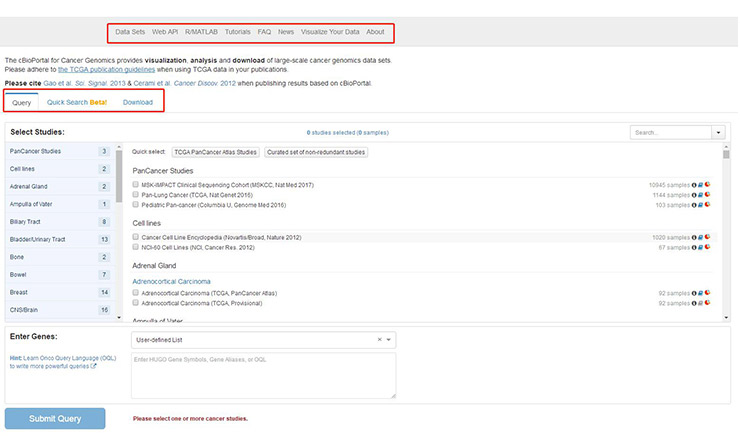
[2] Gao J, Aksoy B A, Dogrusoz U, et al. Integrative Analysis of Complex Cancer Genomics and Clinical Profiles Using the cBioPortal [J]. Science Signaling, 2013, 6(269): pl1-pl1.
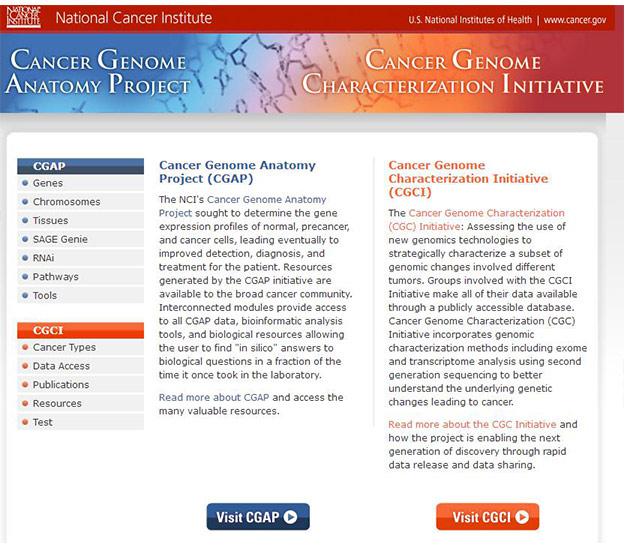
[3] Strausberg R L, Buetow K H, Emmert-Buck M R, et al. The Cancer Genome Anatomy Project: building an annotated gene index [J]. Trends in Genetics Tig, 2000, 16(3): 103-106.
[4] Wilks C, Cline M S, Weiler E, et al. The Cancer Genomics Hub (CGHub): overcoming cancer through the power of torrential data [J]. Database, 2014, 2014: bau093-bau093.
[5] Zhang J, Finney R P, Rowe W, et al. Systematic analysis of genetic alterations in tumors using Cancer Genome WorkBench (CGWB) [J]. Genome Research, 2007, 17(7): 1111-1117.
[6] Forbes S A, Beare D, Gunasekaran P, et al. COSMIC: exploring the world’s knowledge of somatic mutations in human cancer [J]. Nucleic Acids Research, 2015, 43(D1): D805-D811.
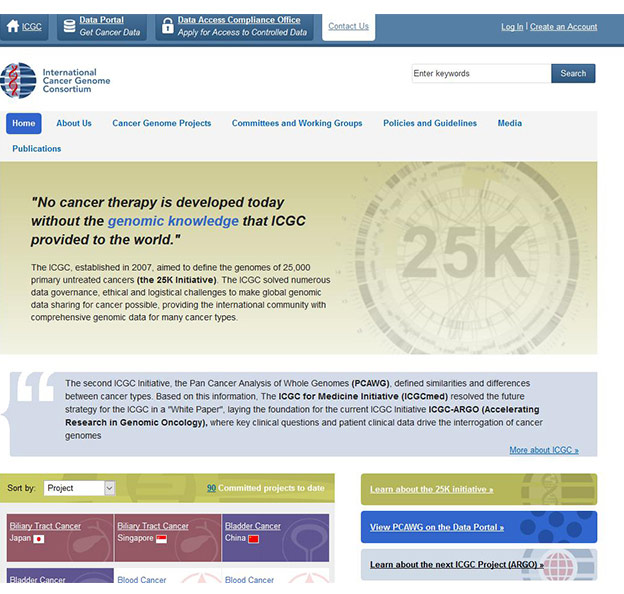
[7] Banks R, LopezOtín, Carlos. International network of cancer genome projects [J]. Nature, 2010, 464(7291): 993-998.
[8] Chang K, Creighton C J, Davis C, et al. The Cancer Genome Atlas Pan-Cancer analysis project [J]. Nature Genetics, 2013, 45(10): 1113-1120.
[9] Benz S C, Craft B, Szeto C, et al. The UCSC Cancer Genomics Browser: update 2011 [J]. Nucleic Acids Research, 2013, 43(Database issue): 812-7.
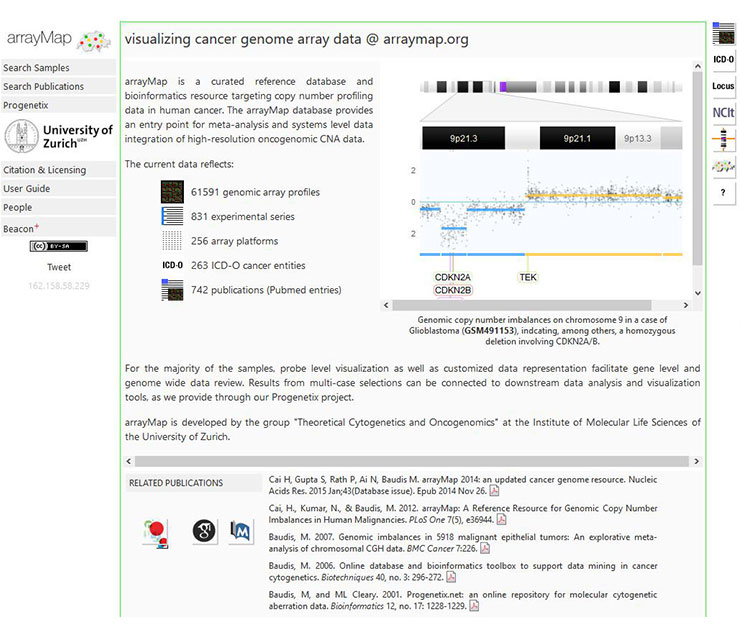
[10] Cai H, Gupta S, Rath P, et al. ArrayMap 2014: An updated cancer genome resource [J]. Nucleic Acids Research, 2014, 43(D1).
[11] Wu T J, Shamsaddini A, Pan Y, et al. A framework for organizing cancer-related variations from existing databases, publications and NGS data using a High-performance Integrated Virtual Environment (HIVE) [J]. Database, 2014, 2014: bau022-bau022.
[12] Scheinin I, Myllykangas S, Borze I, et alet al
[13] Cao Q, Zhou M, Wang X, et al. CaSNP: a database for interrogating copy number alterations of cancer genome from SNP array data [J]. Nucleic Acids Research, 2011, 39(Database issue): D968.
[14] Timms B. Cancer genome project to start [J]. European Journal of Cancer, 2000, 36(6): 687.
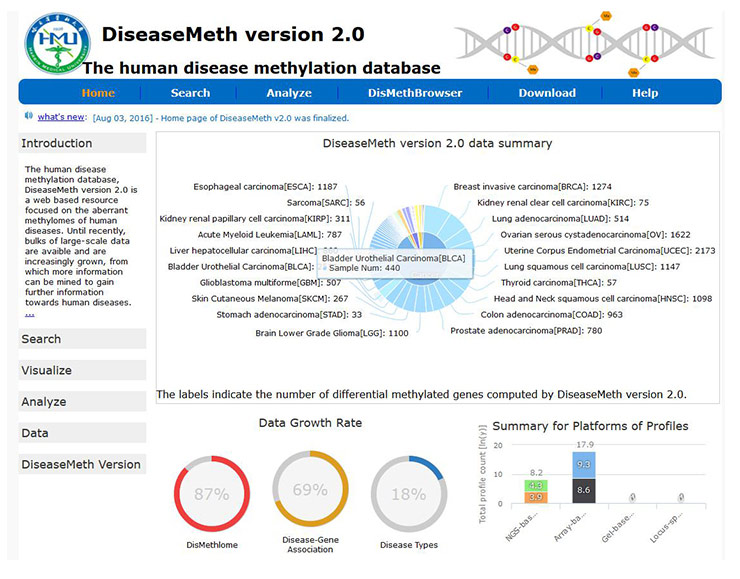
[15] Lv J, Liu H, Su J, et al. DiseaseMeth: a human disease methylation database [J]. Nucleic Acids Research, 2012, 40(Databaseissue): 1030-5.
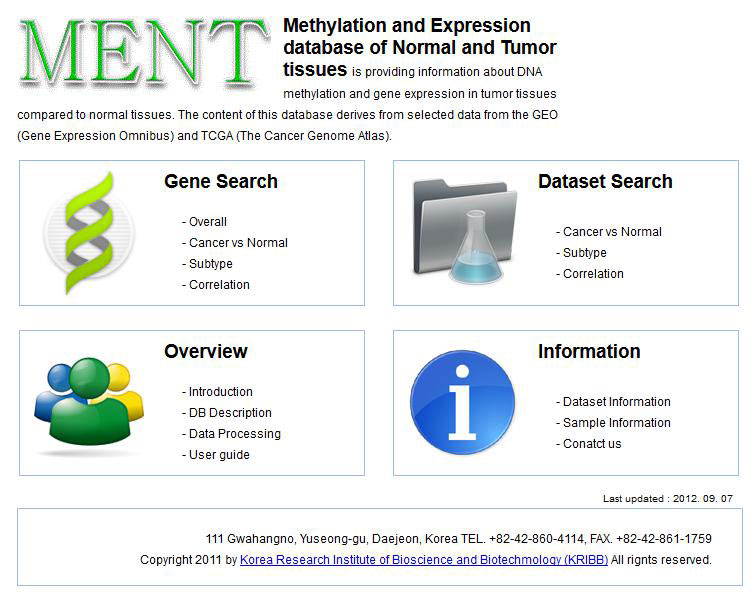
[16] Baek S J, Yang S, Kang T W, et al. MENT: Methylation and expression database of normal and tumor tissues [J]. Gene, 2013, 518(1): 194-200.
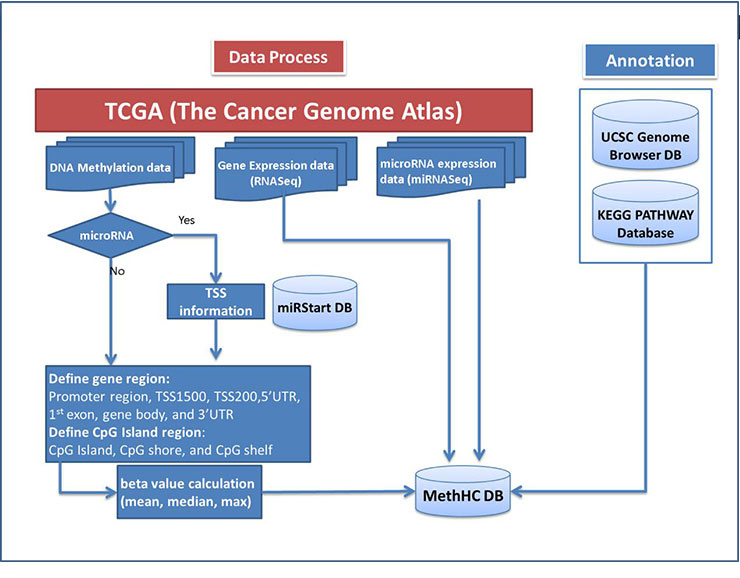
[17] Huang W Y, Hsu S D, Huang H Y, et al. MethHC: a database of DNA methylation and gene expression in human cancer [J]. Nucleic Acids Research, 2015, 43(D1): D856-D861.
[18] He X, Chang S, Zhang J, et al. MethyCancer: the database of human DNA methylation and cancer [J]. Nucleic Acids Research, 2008, 36(Database issue): D836-841.
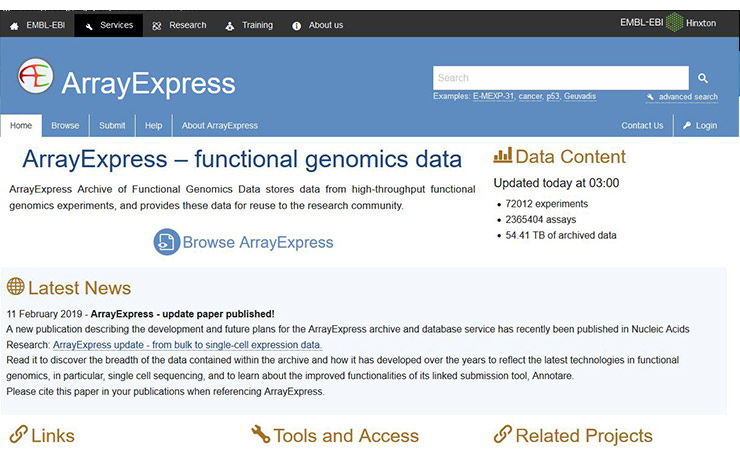
[19] Kolesnikov N, Hastings E, Keays M, et al. ArrayExpress update--simplifying data submissions [J]. Nucleic Acids Research, 2015, 43(D1): D1113-D1116.
[20] Frenkel-Morgenstern M, Gorohovski A, Vucenovic D, et al. ChiTaRS 2.1--an improved database of the chimeric transcripts and RNA-seq data with novel sense-antisense chimeric RNA transcripts [J]. Nucleic Acids Research, 2015, 43(D1): D68-D75.
[21] Barrett T, Troup D B, Wilhite S E, et al. NCBI GEO: archive for functional genomics data sets - 10years on [J]. Nucleic Acids Research, 2012, 39(D1).
[22] Xie B, Ding Q, Han H, et al. miRCancer: a microRNA-cancer association database constructed by text mining on literature [J]. Bioinformatics, 2013, 29(5): 638-644.
[23] Rhodes D R, Kalyana-Sundaram S, Mahavisno V, et al. Oncomine 3.0: Genes, Pathways, and Networks in a Collection of 18,000 Cancer Gene Expression Profiles [J]. Neoplasia, 2007, 9(2): 166-180.
[24] Wang D, Gu J, Wang T, et al. OncomiRDB: a database for the experimentally verified oncogenic and tumor-suppressive microRNAs [J]. Bioinformatics, 2014, 30(15): 2237-2238.
[25] Bhattacharya A, Ziebarth J D, Cui Y. SomamiR: A database for somatic mutations impacting microRNA function in cancer [J]. Nucleic Acids Research, 2012, 41(Database issue).
[26] Porta-Pardo E, Hrabe T, Godzik A. Cancer3D: understanding cancer mutations through protein structures [J]. Nucleic Acids Research, 2015, 43(D1): D968-D973.
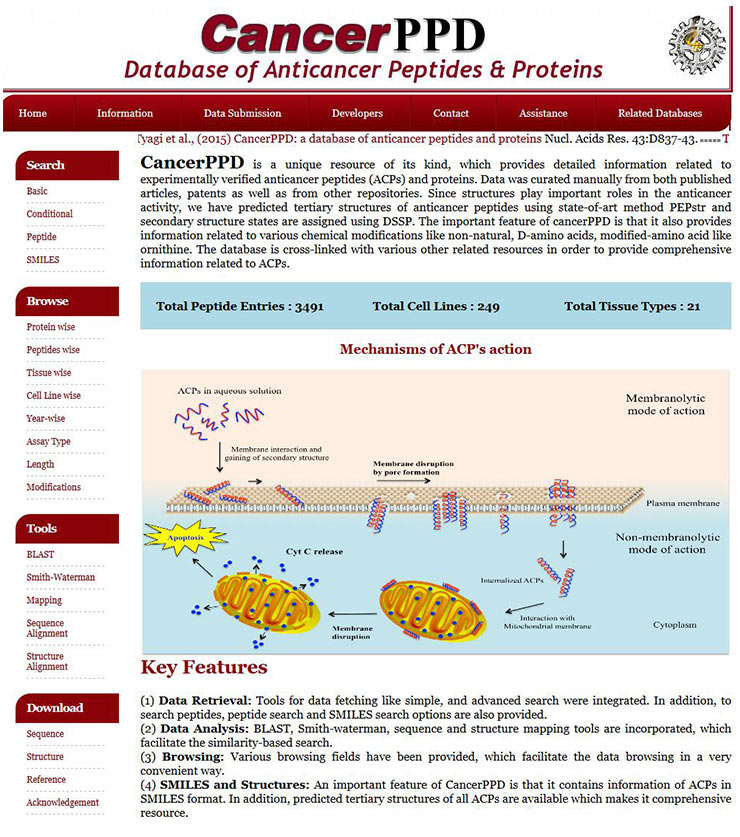
[27] Tyagi A, Tuknait A, Anand P, et al. CancerPPD: a database of anticancer peptides and proteins [J]. Nucleic Acids Research, 2015, 43(D1): D837-D843.
[28] Li J, Duncan D T, Zhang B. CanProVar: a human cancer proteome variation database [J]. Human Mutation, 2010, 31(3): 219-228.
[29] Ellis M J, Gillette M, Carr S A, et al. Connecting genomic alterations to cancer biology with proteomics: The NCI clinical proteomic tumor analysis consortium [J]. Cancer Discovery, 2013, 3(10): 1108-1112.
[30] He Y, Zhang M, Ju Y, et al. dbDEPC 2.0: updated database of differentially expressed proteins in human cancers [J]. Nucleic Acids Research, 2012, 40(D1): D964-D971.
[31] An O, Pendino V, D’Antonio M, et al. NCG 4.0: the network of cancer genes in the era of massive mutational screenings of cancer genomes [J]. Database, 2014, 2014: bau015-bau015.
[32] Leroy B, Fournier J L, Ishioka C, et al. The TP53 website: an integrative resource centre for the TP53 mutation database and TP53 mutant analysis [J]. Nucleic Acids Research, 2013, 41(Database issue): D962.
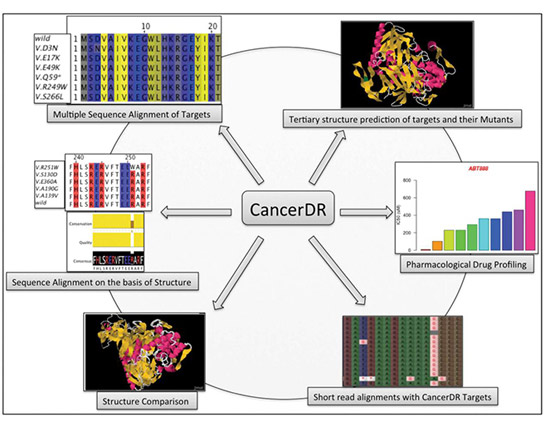
[33] Kumar R, Chaudhary K, Gupta S, et al. CancerDR: Cancer Drug Resistance Database [J]. Scientific Reports, 2013, 3: 1445.
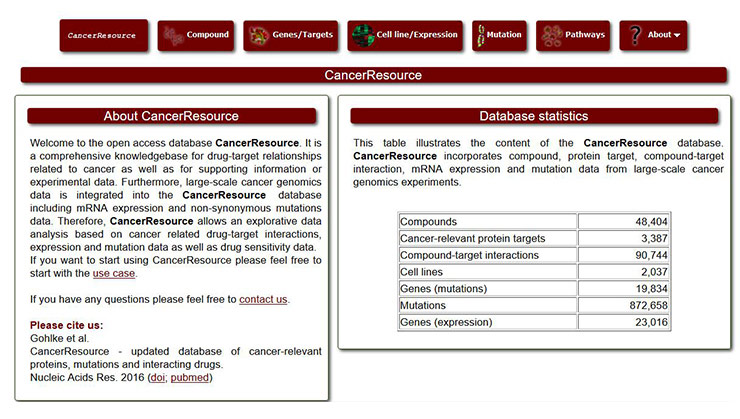
[34] Ahmed J, Meinel T, Dunkel M, et al. CancerResource: a comprehensive database of cancer-relevant proteins and compound interactions supported by experimental knowledge [J]. Nucleic Acids Research, 2011, 39(Database issue): 960-7.
[35] Bulusu K C, Tym J E, Coker E A, et al. canSAR: updated cancer research and drug discovery knowledgebase [J]. Nucleic Acids Research, 2014, 42(D1): D1040-D1047.
[36] Yang W, Soares J, Greninger P, et al. Genomics of Drug Sensitivity in Cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells [J]. Nucleic Acids Research, 2013, 41(Database issue): D955.
[37] Pires D E V, Blundell T L, Ascher D B. Platinum: A database of experimentally measured effects of mutations on structurally defined protein-ligand complexes [J]. Nucleic Acids Research, 2014, 43(D1).








Comments
Leave a Comment