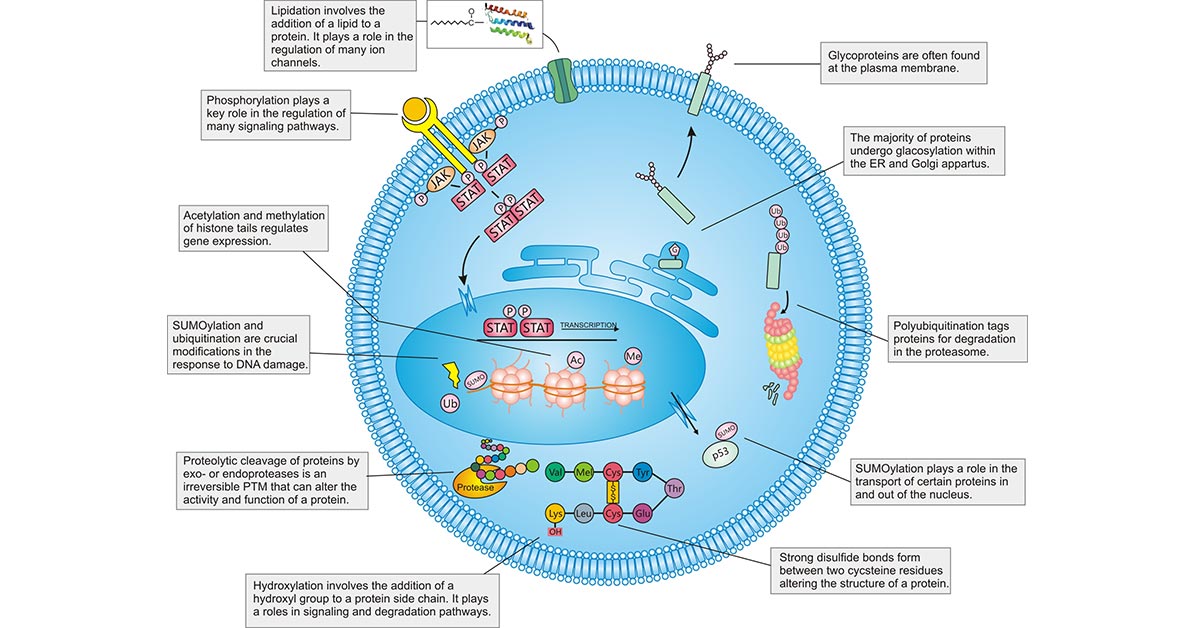
According to the "central dogma" of biology, the DNA is first transcribed to RNA and then translated to proteins. However, most proteins undergo some additional steps after their biosynthesis, which are necessary for cells, tissues, and organisms to achieve their functional biology and diversity. These enzymatic modifications of proteins following protein translation are known as PTMs.
References
[1] Human Genome Sequencing Consortium (2004) Finishing the euchromatic sequence of the human genome. Nature 431: 931–945.
[2] Jensen ON. Modification-specific proteomics: characterization of post-translational modifications by mass spectrometry. Curr Opin Chem Biol 2004, 8: 33–41.
[3] Ramazi, S., Allahverdi, A. and Zahiri, J. Evaluation of post-translational modifications in histone proteins: a review on histone modification defects in developmental and neurological disorders [J]. J. Biosci. 2020, 45, 135.
[4] Blom, N., Sicheritz-Pontén, T., Gupta, R. et al. Prediction of post-translational glycosylation and phosphorylation of proteins from the amino acid sequence [J]. Proteomics, 2004, 4, 1633–1649.
[5] Choudhary, C., Kumar, C., Gnad, F. et al. (2009) Lysine acetylation targets protein complexes and co-regulates major cellular functions. Science, 325, 834–840.
[6] Wellen, KE., Hatzivassiliou, G., et al. ATP-citrate lyase links cellular metabolism to histone acetylation. Science., 2009, 324, 1076–80.
[7] Haltiwanger, R.S. and Lowe, J.B. Role of glycosylation in development. Annu. Rev. Biochem., 2004, 73, 491–537.
[8] Ohtsubo, K. and Marth, J.D. Glycosylation in cellular mechanisms of health and disease. Cell, 2006, 126, 855–867.
[9] Goulabchand, R., Vincent, T., Batteux, F. et al. Impact of autoantibody glycosylation in autoimmune diseases. Autoimmun. Rev., 2014, 13, 742–750.
[10] Han, Z. J., Feng, Y. H., et al. The post-translational modification, SUMOylation, and cancer (Review). Int. J. Oncol.52, 1081–1094 (2018).
[11] Aicart-Ramos, C., Valero, R.A. and Rodriguez-Crespo, I. Protein palmitoylation and subcellular trafficking. Biochim. Biophys. Acta (BBA) Biomembr., 2011, 1808, 2981–2994.
[12] Walsh, C. T., Garneau-Tsodikova, S., and Gatto, G. J. Jr. Protein posttranslational modifications: the chemistry of proteome diversifications. Angew. Chem. Int. Ed. Engl. 2005, 44, 7342–7372.
[13] Mann, M., and Jensen, O. N. Proteomic analysis of post-translational modifications. Nat. Biotechnol. 2003, 21, 255–261.
[14] Ryšlavá, H., Doubnerová, V., Kavan, D. et al. Effect of posttranslational modifications on enzyme function and assembly. J. Proteomics, 2013, 92, 80–109.
[15] Marshall, C. Protein prenylation: a mediator of protein-protein interactions. Science, 1993, 259, 1865–1867.
[16] Haltiwanger, R.S. and Lowe, J.B. Role of glycosylation in development. Annu. Rev. Biochem., 2004, 73, 491–537.
[17] Karve, T.M. and Cheema, A.K. Small changes huge impact: the role of protein posttranslational modifications in cellular homeostasis and disease. J. Amino Acids, 2011, 1–13.
[18] Ohtsubo, K. and Marth, J.D. Glycosylation in cellular mechanisms of health and disease. Cell, 2006, 126, 855–867.
[19] Del Monte, F. and Agnetti, G. Protein post-translational modifications and misfolding: new concepts in heart failure. Proteomics Clin. Appl., 2014, 8, 534–542.
[20] Deribe YL, Pawson T, Dikic I. Post-translational modifications in signal integration. Nat Struct Mol Biol 2010, 17: 666–672.
[21] Zhao S, Xu W, et al. Regulation of cellular metabolism by protein lysine acetylation. Science 2010, 327: 1000–1004.
[22] Nsiah-Sefaa, A. and McKenzie, M. Combined defects in oxidative phosphorylation and fatty acid β-oxidation in mitochondrial disease. Biosci. Rep., 2016, 36, e00313.
[23] Falkenberg, K.J. and Johnstone, R.W. Histone deacetylases and their inhibitors in cancer, neurological diseases and immune disorders. Nat. Rev. Drug Discov., 2014, 13, 673.
[24] Park, G., Tan, J., Garcia, G. et al. Regulation of histone acetylation by autophagy in Parkinson disease. J. Biol. Chem., 2016, 291, 3531–3540.
[25] Micel, L.N., Tentler, J.J., Smith, P.G. et al. Role of ubiquitin ligases and the proteasome in oncogenesis: novel targets for anticancer therapies. J. Clin. Oncol., 2013, 31, 1231.
[26] Popovic, D., Vucic, D. and Dikic, I. Ubiquitination in disease pathogenesis and treatment. Nat. Med., 2014, 20, 1242–1253.
[27] Robertson, K.D. DNA methylation and human disease. Nat. Rev. Genet., 2005, 6, 597.
[28] Sun, G.-D., Cui, W.-P., Guo, Q.-Y. et al. Histone lysine methylation in diabetic nephropathy. J. Diabetes Res., 2014, 1–9.
[29] Lauc, G., Huffman, J.E., Pučić, M. et al. Loci associated with N-glycosylation of human immunoglobulin G show pleiotropy with autoimmune diseases and haematological cancers. PLoS Genet., 2013, 9, e1003225.
[30] Li, S., Li, J., Ning, L. et al. In silico identification of protein S-palmitoylation sites and their involvement in human inherited disease. J. Chem. Inf. Model,2015, 55, 2015–2025.
[31] Meckler, X., Roseman, J., Das, P. et al. Reduced Alzheimer's disease β-amyloid deposition in transgenic mice expressing S-palmitoylation-deficient APH1aL and nicastrin. J. Neurosci., 2010, 30, 16160–16169.
[32] Resh, M.D. Palmitoylation of proteins in cancer. Biochem. Soc. Trans., 2017, 45, 409–416.



Comments
Leave a Comment