This Mouse CXCL10 (IP-10) ELISA Kit provides quantitative measurement of CXCL10 in Mus musculus samples, supporting research into interferon-γ-driven immune signaling and chemokine-mediated T cell recruitment. Because CXCL10 levels can shift dramatically between baseline and inflammatory states, the kit is designed to capture this dynamic range without requiring multiple dilution schemes.
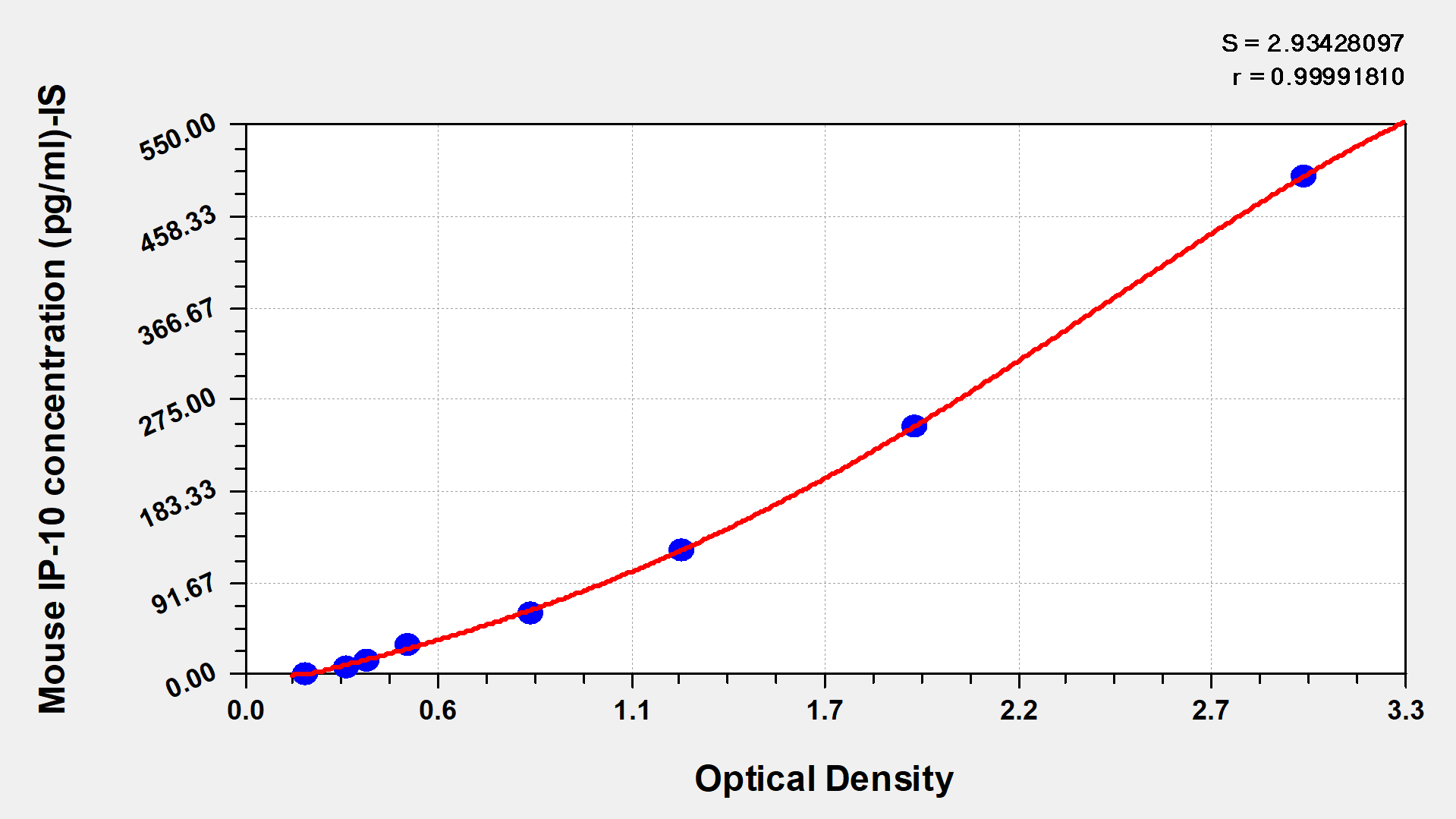
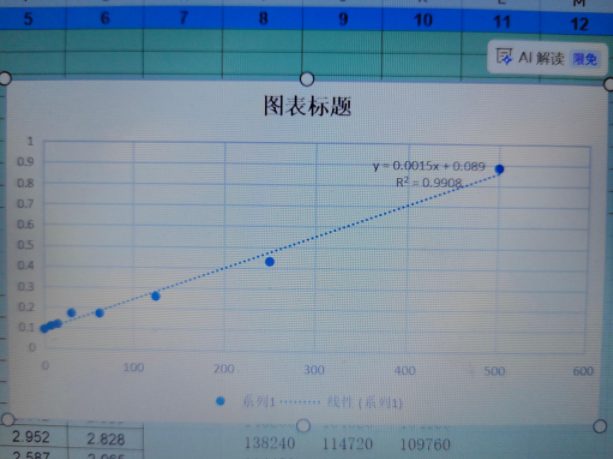
Detection Range: The 7.8–500 pg/mL range spans resting-state serum CXCL10 concentrations in naïve mice (typically 20–80 pg/mL) through the elevated levels seen during viral infection or tumor-driven inflammation (often 200–500+ pg/mL), meaning most samples can be run neat or with minimal dilution.
Sensitivity: At 3.9 pg/mL, the kit detects CXCL10 in low-abundance contexts such as conditioned media from unstimulated splenocytes or early-phase innate immune responses before IFN-γ amplification drives robust chemokine production.
Sample Compatibility: Validation across serum, plasma, and tissue homogenates enables researchers to correlate circulating CXCL10 with local expression in inflamed tissues—critical for dissecting compartmentalized immune responses in organs like lung, liver, or tumor microenvironment.
Assay Efficiency: The 50–100 µL sample volume and 1–5 hour protocol accommodate limited-volume mouse bleeds and same-day processing of tissue panels from multi-timepoint infection or treatment studies.
This kit is well-suited for studies of viral immunopathology, graft rejection, and CXCR3-dependent antitumor immunity.