The winter and spring seasons every year mark the peak period of influenza A virus outbreaks. Due to its numerous hosts in the natural environment and its high potential for mutation or recombination, each outbreak can cause widespread transmission. The influenza A virus has previously caused global concerns and panic, and it still exists in various parts of the world today. As time passes, our understanding of this virus continues to deepen. In this article, we will provide you with the latest scientific knowledge about the influenza A virus, helping you better understand the transmission, symptoms, prevention measures, and the latest treatment methods of this disease.
1. What is influenza A virus?
1.1 Definition of influenza A virus
Influenza viruses are divided into type A, B, C and D. The influenza A virus, also known as influenza A, is highly pathogenic and easily mutated. Almost every worldwide influenza pandemic is caused by the reappearance of new or old subtypes of influenza A virus. Among all subtypes of influenza A virus, H1N1, H5N1 and H7N9 are of particular concern due to their high pathogenicity. The main symptoms after infection with influenza A virus are high fever, cough, nasal discharge, myalgia, and most of them are accompanied by severe pneumonia, which in severe cases leads to death due to multiple organ failure such as heart and kidney.
1.2 Historical background
Throughout history, influenza A virus has caused several influenza pandemics worldwide [1].
● 1918-1920 pandemic
The pandemic caused by the influenza A virus is the most destructive, known as the greatest plague in human history. First discovered in the eastern United States in January 1918, it became popular with the French army in April and then spread rapidly across the globe. The pandemic has infected hundreds of millions of people with an estimated death toll of 50 million to 100 million. According to the serological origin, the pathogen of the pandemic was considered to be porcine Hsw1N1(H1N1) influenza virus.
● 1957-1958 pandemic
The pandemic originated in western Guizhou, China. The H2N2 virus was isolated in Guiyang and spread to Hong Kong in April before spreading to other parts of the world in Southeast Asia and Japan.
● 1968-1969 pandemic
In July 1968, influenza began to appear in Guangdong and Hong Kong regions of China. The virus antigen H3N2 was newly isolated from this influenza and proved to be a mutation of H2N2 virus. Similar to H2N2, H3N2 also has the characteristics of airborne and strong infectious. Most of the infected people in this pandemic are aged over 65.
● 2009-2010 pandemic
The epidemic first broke out in Mexico in March 2009, known as "human swine flu", and then spread rapidly around the world. In April of the same year, the World Health Orgation (WHO) changed its name to "2009 H1N1 influenza".
2. Genome and structure of influenza A virus
Influenza A virus belongs to the Orthomyxoviridae (Orthomyxoviridae). It contains a negative RNA genome encoding 11 essential viral proteins, including:
(1) Hemagglutinin (HA) gene: encoding hemagglutinin surface glycoprotein, the protein is the key for the virus to enter the host cell and also the main antigen of the virus.
(2) Neuraminidase (NA) gene: a surface glycoprotein encode neuraminidase, that protein of which enables the release of the virus from the host cell.
(3)RNA polymerase (PB1, PB1-F2, PB2, PA) gene: subunits encode RNA polymerase that helps that virus replicate its genome in host cells.
(4) Nucleocapsid protein (NP) gene: encoding a nucleocapsid protein, the protein being capable of binding to viral genomic RNA to form a viral nucleocapsid.
(5) Matrix protein (M1, M2) genes: encoding matrix protein, the protein can interact with the nucleocapsid protein to help the virus particle assembly.
(6) Nonstructural protein (NSP1, NSP2) genes: encoding a nonstructural protein, the protein can help virus replication and transmission in host cells.
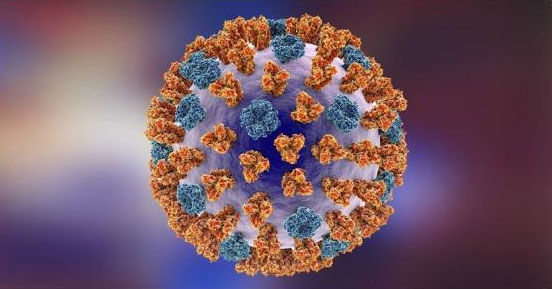
Fig. 1 External structure of influenza A virus
Source: Katryna Kon/Shutterstock.com
The virus has a spherical shell with a diameter of about 80-120 nm. The spherical shell is composed of two major glycoproteins, HA and NA, whose distribution determines the subtype and strain of the virus. Beneath the virus's shell is the nucleocapsid composed of M1, which acts as a protective layer for the inner core and participates in virus replication and assembly. The nucleocapsid contains the genomic fragment of the virus and the protein needed for replication.
3. Epidemiology and transmission of influenza A virus
3.1 Distribution characteristics of influenza A virus
Every winter and spring, influenza A virus causes seasonal influenza outbreaks because of low temperatures and high humidity in winter and spring, an environmental condition conducive to the spread of the virus. In addition, enclosed spaces and crowded places (such as schools, offices, and hospitals) are also important places for the spread of influenza A virus.
In the human population, influenza A virus can cause diseases ranging from mild cold symptoms to severe respiratory infections. Among them, the elderly, young children and those with impaired immune systems are more susceptible to infection and develop severe infections.
3.2 Influenza A virus transmission
Influenza A virus is a highly infectious virus, which is mainly transmitted through the following ways [2–3]:
(1) Respiratory droplet transmission: the influenza A virus can be released into the droplets in the air by the infected person's coughing, sneezing and talking. Other people will be infected by the virus after inhaling the droplets.
(2) Indirect contact transmission: the influenza A virus can survive for several hours after the infected person coughs or sneezes or touches a contaminated surface or object (such as tableware, mobile phone, etc). After other people come into contact with the virus, they can contact with the oral, nasal or ocular mucosa and become infected.
(3) Mother-to-child transmission: pregnant women suffering from influenza A virus may pass the virus to the newborn during childbirth.
In general, the spread of influenza A virus is very diversified, and its high infectious status is one of the main reasons for its large-scale outbreak and epidemic.
4. Pathogenesis and clinical manifestation of influenza A virus
4.1 The mechanism of viral infection and replication
(1) Invasion of host cells: influenza A virus binds to receptors on the surface of host cells through its surface hemagglutinin protein, which is mainly the acidic glycoprotein receptor on the surface of epithelial cells of nose, throat and lung.
(2) Releasing genome: after the virus invades the host cell, its shell is fused with the host cell membrane, and the RNA of the virus genome is released into the host cytoplasm.
(3) Replication genome: RNA of influenza A virus genome is negative-strand RNA, and positive-strand RNA needs to be produced through replication for subsequent virus replication. This process requires the involvement of the viral core protein (NP) as well as RNA polymerase activity.
(4) Synthesis of viral proteins: mRNA in the viral genome guides the host cells to synthesize viral proteins, including hemagglutinin, neuraminidase, and core proteins.
(5) Virus assembly and release: the newly synthesized virus core protein, RNA and membrane protein are assembled into new virus particles in the host cells and released to the outside through the fusion or dissolution mechanism on the host cell membrane to infect other cells or hosts.
4.2 The effect of host immune response on disease
Host immune response is the body's defense mechanism against infected pathogens, including innate immunity and acquired immunity.
For influenza A virus, the innate immune response is the first line of defense against the virus, which can initially clear and defense the virus through inflammatory reactions and other mechanisms. The acquired immune response includes cellular immunity and humoral immunity, which fight against influenza A virus by producing specific antibodies and activating immune cells.
The impact of the intensity and effect of the host immune response on influenza A virus infection is complex. On the one hand, individuals with strong immune responses may be able to eliminate the virus more effectively, thereby alleviating the disease and promoting recovery. On the other hand, overactivated immune response may lead to excessive inflammatory response, triggering serious pathological reactions and tissue damage.
In addition, the intensity and effect of the immune response may be affected by a variety of factors, such as age, health status, immune history, virus strain and vaccination, etc.
4.3 Common clinical features of influenza A infection
Influenza A virus infection usually manifests as a febrile respiratory disease with symptoms including fever, cough, sore throat, body pain and fatigue, and gastrointestinal symptoms including nausea, vomiting and diarrhea in children. Complications of influenza A virus infection include pneumonia, bronchitis, and exacerbation of underlying conditions.
The severity of influenza A virus infection depends on factors such as age, potential health status, and virus subtype.
5. Public health problem and future direction of influenza A virus
5.1 Ongoing research on influenza A virus and its evolution
The ongoing influenza A virus research aims to understand its evolution, pathogenesis and transmission dynamics. Research focuses on monitoring genetic changes in viruses to enable them to evade immune responses and antiviral therapy, as well as identifying new targets for drug development.
5.2 Potential therapies and vaccine innovations
Currently, potential therapies for influenza A virus include the following:
(1) Antiviral drugs: Drug therapy is the most commonly used treatment for influenza A. These drugs alleviate the symptoms and shorten the course of the disease by inhibiting the reproduction of the virus in the body. However, the virus may develop resistance to these drugs. New antiviral drugs are currently being developed. In addition to the above-mentioned neuraminidase inhibitors (oseltamivir and zanamivir), there are also new RNA polymerase inhibitors, such as mavaloxavir, which inhibit virus replication by inhibiting the cap- dependent endonuclease of influenza virus.
(2) Immunotherapy: Immunotherapy refers to fighting against viruses by enhancing the immune system. At present, the vaccine is an effective method to prevent influenza A, which can protect the human body from the virus by activating the immune system to produce antibodies. In addition, researchers are studying the use of monoclonal antibodies to neutralize viruses and the use of interferon and other immunomodulatory drugs to enhance the body's immune response.
(3) Secondary treatment: Secondary treatment includes the use of antipyretic drugs and antitussive drugs to reduce the symptoms of influenza A virus. Additionally, fluid replacement and supportive treatment may also help maintain physical and health conditions.
New vaccines: In addition to traditional flu vaccines, some new vaccines are also being studied, such as vaccines based on protein subunits and vaccines based on DNA or RNA. These new vaccines have higher immunogenicity and better immune effect, and can more effectively prevent the infection of influenza A virus.
6. CUSABIO Provides Products Related to Influenza A Virus for Scientific Research
6.1 Influenza A virus antibody
|
Product name
|
Code
|
Species Reactivity
|
Application
|
|
HA Antibody
|
CSB-PA18939A0Rb
|
Influenza A virus(H7N9)
|
ELISA
|
|
NP Antibody
|
CSB-PA356056LA01IFZ
|
Influenza A virus
|
ELISA
|
|
HA Antibody
|
CSB-PA356317LA01IDG
|
Influenza A virus
|
ELISA
|
|
PA Antibody
|
CSB-PA395880EA01ILR
|
Influenza A virus
|
ELISA
|
|
NP Antibody
|
CSB-PA529597LA01IMP
|
Influenza A virus
|
ELISA
|
|
HA Antibody
|
CSB-PA607773LA01IER
|
Influenza A virus
|
ELISA, WB
|
|
NP Antibody
|
CSB-PA325769LA01IIM
|
Influenza A virus
|
ELISA, WB
|
|
NS Antibody
|
CSB-PA356065LA01IIE
|
Influenza A virus
|
ELISA
|
|
NS Antibody
|
CSB-PA604972LA01IER
|
Influenza A virus
|
ELISA, WB
|
6.2 Influenza A virus recombinant protein
|
Product name
|
Code
|
Express System
|
|
Recombinant Influenza A virus Hemagglutinin(HA),partial
|
CSB-RP189374BA
|
E.coli
|
|
Recombinant Influenza A virus Hemagglutinin(HA),partial
|
CSB-EP607773IER
|
E.coli
|
|
Recombinant Influenza A virus Hemagglutinin(HA)
|
CSB-EP356317IDG
|
E.coli
|
|
Recombinant Influenza A virus Hemagglutinin(HA) ,partial
|
CSB-EP714917IEX
|
E.coli
|
|
Recombinant Influenza A virus Hemagglutinin(HA) ,partial
|
CSB-EP721113IFG
|
E.coli
|
|
Recombinant Influenza A virus Hemagglutinin(HA),partial
|
CSB-EP3563GMC
|
E.coli
|
|
Recombinant Influenza A virus Hemagglutinin(HA)(X347S,X348S,X509S,X538S),partial
|
CSB-YP356048IBA
|
Yeast
|
|
Recombinant Influenza A virus Hemagglutinin(HA)
|
CSB-YP356317IDG
|
Yeast
|
|
Recombinant Influenza A virus Hemagglutinin(HA),partial
|
CSB-MP3563GMC
|
Mammalian cell
|
|
Recombinant Influenza A virus Hemagglutinin(HA),partial
|
CSB-MP3563GMC1
|
Mammalian cell
|
|
Recombinant Influenza A virus Hemagglutinin(HA),partial
|
CSB-MP3563GMCd7
|
Mammalian cell
|
|
Recombinant Influenza A virus Matrix protein 2(M),partial
|
CSB-EP389902ILU1
|
E.coli
|
|
Recombinant Influenza A virus Matrix protein 1(M)
|
CSB-EP389592ILS
|
E.coli
|
|
Recombinant Influenza A virus Matrix protein 2(M),partial
|
CSB-EP356944IFZ1
|
E.coli
|
|
Recombinant Influenza A virus Matrix protein 2(M)
|
CSB-CF389902ILU
|
in vitro E.coli expression system
|
|
Recombinant Influenza A virus Matrix protein 2(M)
|
CSB-CF356944IFZ
|
in vitro E.coli expression system
|
|
Recombinant Influenza A virus Matrix protein 2(M)
|
CSB-CF389902ILUb0
|
in vitro E.coli expression system
|
|
Recombinant Influenza A virus Matrix protein 1(M1),partial
|
CSB-EP3562GMC1
|
E.coli
|
|
Recombinant Influenza A virus Matrix protein 2(M2)
|
CSB-CF3574GMC
|
in vitro E.coli expression system
|
|
Recombinant Influenza A virus Neuraminidase(NA),partial
|
CSB-EP526039IMR
|
E.coli
|
|
Recombinant Influenza A virus Nuclear export protein(NEP)
|
CSB-EP3575GMC
|
E.coli
|
|
Recombinant Influenza A virus Nucleoprotein(NP)
|
CSB-EP304802IMB
|
E.coli
|
|
Recombinant Influenza A virus Nucleoprotein(NP)
|
CSB-EP325878IMI
|
E.coli
|
|
Recombinant Influenza A virus Nucleoprotein(NP)
|
CSB-EP529597IMP
|
E.coli
|
|
Recombinant Influenza A virus Nucleoprotein(NP)
|
CSB-EP325769IIM
|
E.coli
|
|
Recombinant Influenza A virus Nucleoprotein(NP)
|
CSB-EP630945IFY
|
E.coli
|
|
Recombinant Influenza A virus Nucleoprotein(NP)
|
CSB-EP324412IES
|
E.coli
|
|
Recombinant Influenza A virus Nucleoprotein(NP)
|
CSB-EP323566IEC
|
E.coli
|
|
Recombinant Influenza A virus Nucleoprotein(NP)
|
CSB-EP302450IGF
|
E.coli
|
|
Recombinant Influenza A virus Nucleoprotein(NP)
|
CSB-EP326264IFQ
|
E.coli
|
|
Recombinant Influenza A virus Nucleoprotein(NP)
|
CSB-EP333131IEB
|
E.coli
|
|
Recombinant Influenza A virus Nucleoprotein(NP)
|
CSB-EP356056IFZ
|
E.coli
|
|
Recombinant Influenza A virus Nucleoprotein(NP)(X477S)
|
CSB-EP3561GMC
|
E.coli
|
|
Recombinant Influenza A virus Nucleoprotein(NP)
|
CSB-YP356056IFZ
|
Yeast
|
|
Recombinant Influenza A virus Nucleoprotein(NP)
|
CSB-YP325878IMI
|
Yeast
|
|
Recombinant Influenza A virus Non-structural protein 1(NS)
|
CSB-EP604972IER
|
E.coli
|
|
Recombinant Influenza A virus Non-structural protein 1(NS)
|
CSB-EP391362ILO
|
E.coli
|
|
Recombinant Influenza A virus Non-structural protein 1(NS)
|
CSB-EP361072IAX
|
E.coli
|
|
Recombinant Influenza A virus Non-structural protein 1(NS)
|
CSB-YP604972IER
|
Yeast
|
|
Recombinant Influenza A virus Polymerase acidic protein(PA),partial
|
CSB-EP881268IIO
|
E.coli
|
|
Recombinant Influenza A virus Polymerase acidic protein(PA),partial
|
CSB-YP395880ILR1
|
Yeast
|
|
Recombinant Influenza A virus protein PB1-F2(PB1)
|
CSB-EP455303IMU
|
E.coli
|
|
Recombinant Influenza A virus Protein PB1-F2(PB1)
|
CSB-EP313650IGJ
|
E.coli
|
6.3 Influenza A virus ELISA kit (containing influenza B virus)
|
Product name
|
Code
|
Size
|
|
Human anti-parainfluenza virus(PIV)antibody(IgG) ELISA kit
|
CSB-E09022h
|
96T,5×96T,10×96T
|
|
Human anti-parainfluenza virus(PIV) antibody(IgM) ELISA Kit
|
CSB-E09530h
|
96T,5×96T,10×96T
|
|
Human Influenza B virus (FluB) antibody(IgM) ELISA kit
|
CSB-EQ027333HU
|
96T,5×96T,10×96T
|
|
Human Influenza A virus (FluA) antibody(IgM) ELISA kit
|
CSB-EQ027335HU
|
96T,5×96T,10×96T
|
|
Human Influenza A virus antibody(IgG) ELISA Kit
|
CSB-E14083h
|
96T,5×96T,10×96T
|
|
Human influenza B virus (FluB) antibody (IgG) ELISA Kit
|
CSB-E14084h
|
96T,5×96T,10×96T
|
References
[1]Jeffery K T, John C K. Influenza Virus Evolution, Host Adaptation, and Pandemic Formation[J]. Cell Host & Microbe, 2010, 7(6): 440-451.
[2]https://www.who.int/news-room/fact-sheets/detail/influenza-(avian-and-other-zoonotic)
[3]https://www.cdc.gov/flu/about/disease/spread.htm
[4]https://www.cdc.gov/flu/treatment/whatyoushould.htm
CUSABIO team. Exploring the Science of Influenza A Virus: What You Need to Know. https://www.cusabio.com/c-21108.html



Comments
Leave a Comment