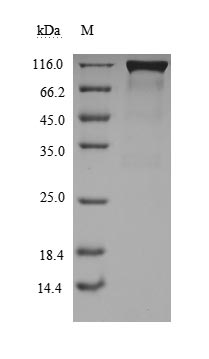
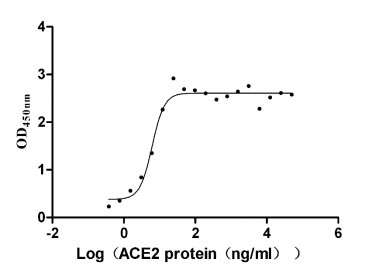
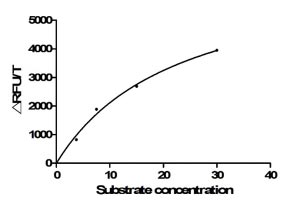
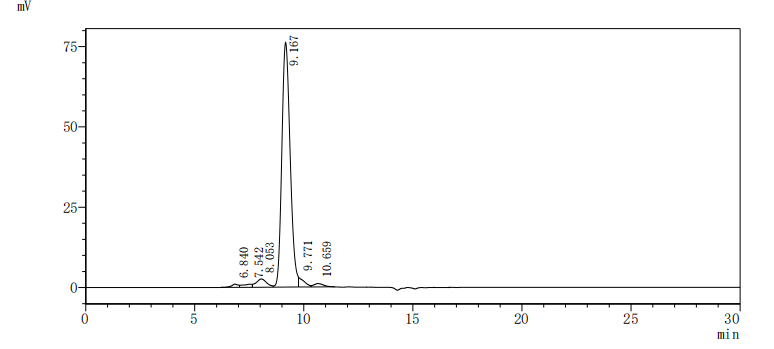
ACE2 is a type I membrane protein that functions as the primary receptor for SARS-CoV-2 spike protein, catalyzing the conversion of angiotensin II to angiotensin-(1-7) and playing a critical role in cardiovascular and immune regulation, thus precise characterization of its binding and enzymatic activity is essential for viral entry studies and therapeutic development. This recombinant Paguma larvata ACE2 covers the extracellular region from amino acid 18 to 740 and is produced in mammalian cells with a C-terminal hFc1 tag, preserving native conformation and post-translational modifications necessary for functional integrity. Functional ELISA demonstrates that immobilized SARS-CoV S-RBD (CSB-MP348663HQE) at 2 μg/ml binds Paguma larvata ACE2 with an EC50 of 5.056–7.559 ng/ml, confirming specific receptor-ligand interaction; enzyme activity is measured via cleavage of the fluorogenic peptide substrate Mca-Ala-Pro-Lys(Dnp)-OH, yielding a Km of 22.84 μM. Purity exceeds 90% as determined by both SDS-PAGE and SEC-HPLC, and endotoxin levels are below 1.0 EU/μg, satisfying stringent requirements for cell-based assays and structural biology applications. This active, highly pure, and endotoxin-controlled reagent supports use in viral entry inhibition studies, receptor-binding kinetics, competitive binding assays, and antibody epitope mapping targeting Paguma larvata ACE2.