What is Oxidative Phosphorylation?
Oxidative phosphorylation, also known as electron transport-linked phosphorylation, refers to the metabolic pathway in which the energy released by nutrients during oxidation is utilized to generate ATP through electrical transport chain. And it is an important cellular energy conversion process and the final process of cell respiration in eukaryotes.
Oxidative phosphorylation occurs in the mitochondrial inner membrane of eukaryotic cells or the cytoplasm of prokaryotes.
Oxidative Phosphorylation & Substrate-Level Phosphorylation
When it comes to oxidative phosphorylation, we have to talk about its "good partner"--substrate-level phosphorylation.
Substrate-level phosphorylation is a metabolic reaction in which the energy-rich phosphorylated compound resulting from the coupled reaction transfers its phosphate group to ADP for ATP synthesis. Or GDP is recharged a phosphate group to generate GTP.
● The Similarities between Oxidative Phosphorylation and Substrate-level Phosphorylation
The main similarity between oxidative phosphorylation and substrate-level phosphorylation is that both their ultimate production is ATP.
● The Differences between Oxidative Phosphorylation and Substrate-Level Phosphorylation
The biggest difference between oxidative phosphorylation and substrate-level phosphorylation is the source of the energy needed to convert ADP to ATP. Substrate level phosphorylation directly phosphorylates ADP to ATP by using the energy from a coupled reaction. While oxidative phosphorylation involves two coupled reactions that are considered to simultaneously occur. In the period of oxidative phosphorylation, the energy produced during the oxidative reaction is transferred to ADP to form ATP.
The Function of Oxidative Phosphorylation
Oxidative phosphorylation provides bulk ATP for living organisms, and the ATP is the main energy source for maintaining life activity. Oxidative phosphorylation also involves the formation of reactive oxygen species (ROS) and the regulation of apoptosis.
The Process of Oxidative Phosphorylation
When a hydroelectric dam works, it converts potential energy released from the falling water into kinetic energy, which turns into electrical energy. Similar to the steps of generating electricity from a hydroelectric dam, ADP makes ATP by a process called chemiosmosis during oxidative phosphorylation.
In eukaryotes, when catabolism such as glycolysis or citric acid cycle occurs, NADH is produced, which is a coenzyme containing a very high transfer electrical potential. When NADH is oxidized in the mitochondrial matrix, its electrons pass through the electron transport chain (ETC) to the electron receptor-oxygen, and simultaneously releases energy that pumps the resulting hydrogen ions through the inner mitochondrial membrane. It spontaneously forms an electrochemical concentration gradient across the inner mitochondrial membrane due to a higher concentration of hydrogen ions in the intermembrane space and a lower concentration in the matrix. When hydrogen ions pass through the inner mitochondrial membrane across electrochemical gradient, ATP synthase captures the proton-motive force for the production of ATP. This process is called chemiosmosis.
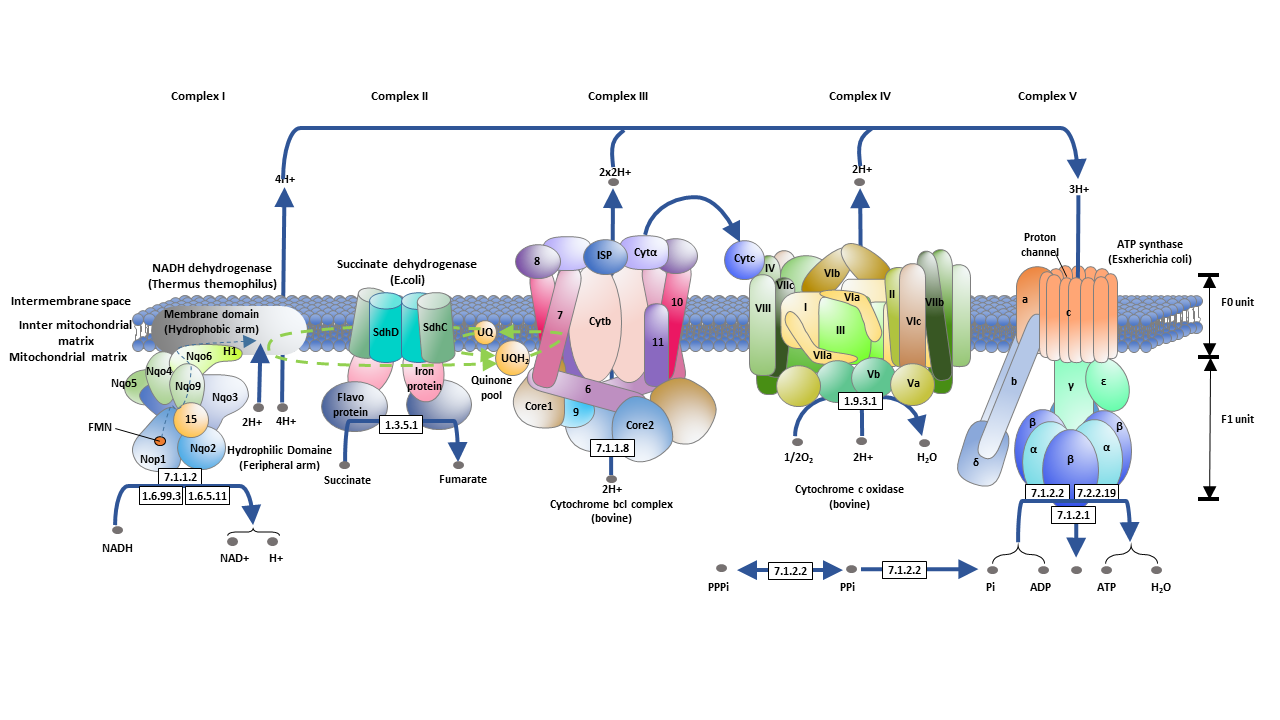
The electron transport chain is a series of proteins located on the inner membrane of the mitochondria.
● NADH-Coenzyme Q Oxidoreductase
The first enzyme in the electron transport chain is the NADH-CoQ oxidoreductase, also known as NADH dehydrogenase or complex I, which is the first entry of protons through the electron transport chain. It catalyzes the oxidation of NADH through coenzyme Q10. As two electrons pass through complex I, four protons are pumped from the mitochondrial matrix into the intermembrane space.
● Succinic-Coenzyme Q Oxidoreductase
The second enzyme that allows protons to passes through the electron transport chain is succinic-coenzyme Q oxidoreductase, also known as succinate dehydrogenase or complex II. It catalyzes the oxidation of succinic acid to form fumarate and the reduction of coenzyme Q10 to ubiquinone (QH2). This reaction does not involve the transfer of electrons, nor does it pump out protons, providing less energy to compare with the oxidation process of NADH. The third entry to the proton on the electron transport chain is electron transfer flavin-coenzyme Q oxidoreductase, also known as electron transfer flavin dehydrogenase, which reduces Q10 by using electrons from electron transfer flavin in the mitochondrial matrix.
● Coenzyme Q-cytochrome C Reductase
Coenzyme Q-cytochrome C reductase, also known as complex III, catalyzes the oxidation of QH2, and the reduction of cytochrome c and ferritin. In this reaction, cytochrome C carries an electron. Coenzyme Q is reduced to QH2 on one side of the mitochondrial membrane, while QH2 is oxidized to coenzyme Q10 on the other side, resulting in the transfer of protons on the membrane, which also contributes to the formation of proton gradients.
● Cytochrome c Oxidase
The last protein complex in the electron transport chain is cytochrome c oxidase, also called complex IV. It mediates the final reaction on the electron transport chain - transferring electrons to the final electron receptor oxygen - oxygen reduces to water - pumping protons through the membrane. At the end of this reaction, protons that directly pumped out and that consumed by the reduction of oxygen to water increase the proton gradient.
Finally, the proton-motive force generated by the proton concentration gradient drives the ATP synthase to phosphorylate ADP to form ATP.
There is another electron-donating molecule - FADH2 in eukaryotes. FADH2 is also the intermediate metabolite during the earlier stage of cellular respiration such as glycolysis or citric acid cycle. In the FADH2 electrical transport chain, FADH2 bypasses the complex I and enters the electrical transport chain by the complex II because it contains less electrical potential than NADH. FADH2 is oxidized to FAD and coenzyme Q is reduced to QH2 in the reaction. And this reaction does not pump out protons either. The subsequent reactions are nearly the same as those in the NADH2 electron transport chain.
Prokaryotes such as bacteria and archaea have many electron transfer enzymes that can use a very wide range of chemicals as substrates. As the same with eukaryotes, electron transport in prokaryotic cells also uses the energy released by oxidation from the substrate to pump protons across the membrane to create an electrochemical gradient, which drives ATP synthase to generate ATP. The difference is that bacteria and archaea use many different substrates as electron donors or electron receptors. This also helps prokaryotes to survive and grow in different environments.
Factors Affecting Oxidative Phosphorylation
● Inhibitors
Under normal conditions, electron transfer and phosphorylation are tightly coupled. Some compounds can affect electron transport or interfere with phosphorylation reactions, all of which cause oxidative phosphorylation abnormalities. Here introduce four factors affecting oxidative phosphorylation.
Respiratory chain inhibitor: A substance that blocks electron transport at a certain part of the respiratory chain and inhibits the oxidation process. Some respiratory chain inhibitors bind to iron-sulfur proteins in NADH-Q reductase and block the transmission of electrons from NADH to CoQ, such as rotenone, phenoxymycin A, and barbital, ampicillin. Some substances inhibit the electron transfer between Cytb and Cytc1, such as antimycin A and dimercaptopropanol. Cyanide, azide, H2S, and C0 inhibit cytochrome oxidase, making electrons unable to pass to oxygen.
Oxidative phosphorylation inhibitors: These reagents directly interfere with the formation of ATP and also prevent electron transfer. The combination of oligomycin and dicyclohexylcarbonyldiimide with the F0 unit of ATP synthase prevents the hydrogen ions from flowing back from the proton channel, rendering the phosphorylation process incomplete, thus blocking the oxidative phosphorylation of intact mitochondria.
Uncoupling agent: The uncoupling agent separates the two coupling processes of electron transfer and ATP synthesis. Such compounds only inhibit the formation of ATP, but do not affect the electron transfer process. So the free energy generated by electron transfer is converted into heat energy, which excessively uses oxygen and fuel substrates. Such agents cause the electron transfer to lose normal control, resulting in excessive utilization of oxygen and fuel substrates, and energy is not stored. A typical uncoupler is 2,4-dinitrophenol (DNP). Because DNP is a fat-soluble substance, it can move freely in the mitochondrial membrane. When it enters the matrix, it can release H+. Return to the cytosol side. The H+ can be combined to eliminate the transmembrane gradient of H+, so that the energy released by the oxidation process cannot be used for the synthesis reaction of ATP, but referred to as a proton carrier.
● The Regulation of ADP
The rate of oxidative phosphorylation in normal organisms is mainly regulated by ADP. When the body uses ATP increase, the ADP concentration increases and the oxidative phosphorylation rate is increased after transporting into the mitochondria; otherwise, the ADP deficiency causes the oxidative phosphorylation rate to slow down. This regulation allows the rate of ATP production to adapt to physiological needs.
● Thyroid Hormone
Thyroid hormone can activate Na+-K+ATPase on the cell membrane of many tissues, accelerate the decomposition of ATP into ADP and Pi, and increase the number of ADP into mitochondria, thus decreasing the ATP/ADP ratio and accelerating the oxidative phosphorylation rate. As the synthesis and decomposition rate of ATP increases, the body's oxygen consumption and heat production increase, the basal metabolic rate increases, and the basal metabolic rate are one of the most important clinical indications for patients with hyperthyroidism.
● Mitochondrial DNA Mutation
Due to its naked circular double helix structure and the absence of protein protection and damage repair system of mitochondrial DNA (mtDNA), it is susceptible to mutate by oxidative phosphorylation. mtDNA encodes 13 proteins involved in oxidative phosphorylation. Therefore, mtDNA mutations can affect the oxidative phosphorylation process, resulting in a decrease in ATP yield and thus leading to many related diseases.
Diseases and Abnormal Oxidative Phosphorylation
Oxidative phosphorylation exerts a multiply role in the body. So once it is abnormal, it will cause diseases.
Many mitochondrial diseases are linked to defective oxidative phosphorylation. And tissues with high energy requirements are particularly susceptible to undergo oxidative phosphorylation defects, like brain, nerves, retina, bone and heart muscle. When there is impairment in oxidative phosphorylation in these tissues, it could clinically manifest as seizures, hypotonia, ophthalmoplegia, convulsions, muscle weakness, and cardiomyopathy, etc.



