With the continuous improvement of living standards, obesity has become one of the major problems troubling many people. Losing weight is a lifetime compulsory course for many people. A finding found by H. S. Park et al. showed that PPARγ NEDD8-based neddylation essential for adipogenesis is a potential target for treating obesity [1].
This article will mainly introduce the definition and structure of NEDD8, NEDD8-based neddylation pathway and function, as well as NEDD8' roles in different diseases.
1. NEDD8 Structure
In 1992, Kumar et al. first discovered neural precursor cell expressed, developmentally down-regulated 8 (NEDD8) using a subtraction cloning method [2]. NEDD8 is a small ubiquitin-like (UBL) modifier covalently attached to certain lysine residues of a target substrate protein, a process known as neddylation that is very similar to ubiquitination.
As the ubiquitin's closest relative, NEDD8 shares about 60% sequence identity and 80% sequence homology with ubiquitin [3]. Therefore, NEDD8 still retains many of the structural characteristics that determine ubiquitin's ability to label proteins for subsequent degradation.
Structurally, NEDD8 is composed of a globular core known as the β-grasp fold that contains four five-stranded mixed β-sheets featured by an α-helix and two 310 helices, a flexible C-terminal tail that extends out from the globular domain body and ends with the Gly75-Gly76 motif, and a ubiquitin-fold domain (UFD) [3][4]. Mutational analysis has shown that Gly76 is required for the conjugation of NEDD8 to substrate proteins.
Like ubiquitin, NEDD8 displays an almost identical arrangement of acidic and hydrophobic surfaces centered on Ile44, also known as an Ile44-centered hydrophobic patch, including residues of Leu8, Ile44, His68, and Val70.
Majority of the amino acid sequence differences between NEDD8 and ubiquitin map to the surface. And ample studies have clearly shown that these small but substantial amino acid differences guarantee unambiguous distinction between substrates directed to ubiquitination or neddylation.
The neddylation substrates are mainly divided into two categories: best-characterized cullins and less-studied non-cullin substrates. Cullins are scaffold components of the cullin-RING ligases (CRLs), the largest family of E3 ubiquitin ligases, which are composed of 8 cullin members, adaptor proteins, substrate recognition receptors, and the RING family members.
Table 1. Components of cullin-RING ligases (CRLs)
2. NEDD8 Pathway and Function
NEDD8 uses similar enzymatic machinery for its conjugation and deconjugation as ubiquitin. The NEDD8 conjugation cascade, also known as neddylation, is a multi-enzymatic event involving E1 NEDD8-activating enzyme (NAE), E2 NEDD8-conjugating enzyme, E3 NEDD8 ligase, and de-neddylation enzymes.
NEDD8 is first synthesized as a precursor and then converted into the mature form after undergoing a proteolytic process that exposes its C-terminal Gly-Gly residues. The Gly-Gly motif serves as the attachment site for target substrates [5]. This process is catalyzed by UCHL3 and NEDP1/DEN1/SENP8 [6][7].
The mature NEDD8 is adenylated and activated by NEDD8-activating enzyme E1 (NAE) in a Mg2+/ATP-dependent manner [8]. This reaction generates the E1-NEDD8 complex with the formation of a thioester linkage through NEDD8's C-terminal Gly-76 residue with the catalytic subunit UBA3 of E1.
The NEDD8 E1 is a heterodimer made up of N-terminal APPBP1/NAE1 and C-terminal UBA3 [9]. NAE1 consists of three domains: an adenylation domain, a catalytic cysteine-containing domain that is involved in the E1-NEDD8 thioester linkage, and a ubiquitin-fold domain (UFD) that binds to the E2 [10].
Activated NEDD8 is subsequently transferred to the catalytic cysteine in UBA3 to form E1-NEDD8 [11], and the NEDD8-AMP complexed to E1 is formed concomitantly during the transferring process.
This E1-NEDD8-AMP ternary complex delivers one molecule of NEDD8 to the cysteine active site of the NEDD8-conjugating enzyme E2, either UBC12 or UBE2F, via transthiolation reaction.
Finally, a NEDD8 ligase E3-carrying substrate binds to E2-activated NEDD8 and then transfers NEDD8 over to the ε-amino group of the lysine residue in the substrate protein to form an isopeptide linkage. NEDD8 can be released from the NEDD8 isopeptide complex under the action of deneddylases such as COP9 signaling subunit 5 and ubiquitin-degrading enzyme 1, thus entering the next NEDD8-conjugation pathway cycle.
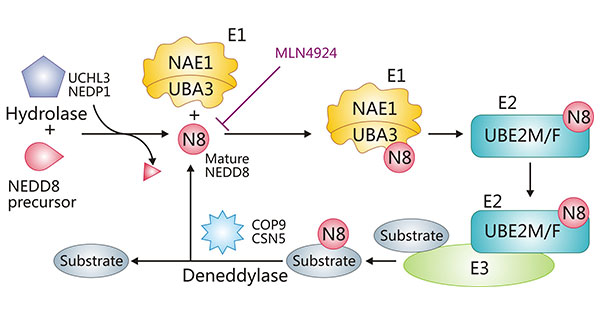
Figure 1. The schematic representation of NEDD8 activation, conjugation, and deneddylation
NEDD8 acts as the on-switch of about one-third of all E3 ubiquitin ligases. The NEDD8 conjugation pathway is essential for in cellular function through its prominent role in mediating the ubiquitination by CRLs of numerous substrate proteins especially cullins that are involved in cell cycle regulation, transcription, signal transduction, cell proliferation, apoptosis, DNA repair, and tumorigenesis.
3. NEDD8 and Disease Research
Protein neddylation is involved in various physiological and pathological events, such as metabolic processes including adipogenesis, lipid droplet formation, redox homeostasis, some neurodegenerative diseases, and cancer. Dysregulated neddylation has been observed in tumors and neurodegenerative diseases.
3.1 NEDD8 in adipogenesis
H-S Park et al. demonstrated that the NEDD8 -based neddylation of PPARγ plays an essential role in adipogenesis [1]. During adipogenesis, NEDD8 is highly expressed in preadipocytes and conjugates with PPARγ, resulting in PPARγ stabilization. When blocking the neddylation process by NEDD8-targeting siRNAs (or viral vectors) or a selective inhibitor of E1, MLN4924, adipocyte differentiation and fat tissue development were severely hampered. This finding suggests that neddylation could be a new therapeutic target for obesity and obesity-related diseases.
3.2 NEDD8 in cancer treatment
Neddylation, a novel post-translation modification of proteins, is over-activated in a variety of tumors and is closely related to the occurrence and development of cancer. Inhibition of the overexpressed NEDD8-based neddylation pathway members may serve as a new anticancer strategy.
Studies have shown that NEDD8 is highly expressed in hepatocellular carcinoma, and its expression level is negatively correlated with the prognosis of patients. The mRNA and protein expression levels of NEDD8 were significantly up-regulated in esophageal squamous cell carcinoma, and the overall survival rate of patients was low.
Both in vivo and in vitro experiments showed that down-regulated NEDD8 could significantly inhibit tumor growth. Down-regulation of NEDD8 induces DNA damage and activates the death receptor 5 (DR5) -dependent exogenous apoptotic pathway, thereby killing tumors.
In addition, the knockdown of NEDD8 inactivated CRL, resulting in its substrate cyclin-dependent kinase inhibitor 1A, p21, cyclin-dependent kinase inhibitor 1B (p21), and G2 checkpoint kinase Wee1 accumulated in large amounts, thus inducing cell cycle arrest in the G2 phase.
An overall increase in the level of NEDD8 conjugation has been detected in oral squamous cell carcinoma cell lines where NEDD8 pathway blockage reduced cell proliferation [12].
Pharmaceutical agents can be used to affect the NEDD8 pathway. MLN4924 is a suicide inhibitor of NEDD8 E1 and inhibits tumor cell proliferation. It is undergoing a phase II clinical trial for the treatment of acute myelocytic leukemia and could be the first drug targeted at UBL in the treatment of human cancer.
CUSABIO provides many proteins that are involved in the NEDD8-based neddylation for associated studies. These proteins are only used for scientific research.
References
[1] Park, HS., Ju, UI., Park, JW. et al. PPARγ neddylation essential for adipogenesis is a potential target for treating obesity [J]. Cell Death Differ 23, 1296–1311 (2016).
[2] Kumar, S. , Tomooka, Y. , et al. (1992) Identification of a set of genes with developmentally down-regulated expression in the mouse brain [J]. Biochem. Biophys. Res. Commun.. 185, 1155–1161.
[3] Kamitani T, Kito K, Nguyen HP, Yeh ET. Characterization of NEDD8, a developmentally down-regulated ubiquitin-like protein [J]. J Biol Chem. 1997;272:28557–62.
[4] Whitby FG, Xia G, et al. Crystal structure of the human ubiquitin-like protein NEDD8 and interactions with ubiquitin pathway enzymes [J]. The Journal of Biological Chemistry. 1998;273:34983-34991.
[5] T. Kamitani, K. Kito, et al. Characterization of NEDD8, a developmentally down-regulated ubiquitin-like protein [J]. J. Biol. Chem., 272 (1997), pp. 28557-28562.
[6] Mendoza HM, Shen LN, et al. (2003) NEDP1, a highly conserved cysteine protease that deNEDDylates Cullins [J]. J Biol Chem 278: 25637–25643.
[7] Wu K, Yamoah K, et al. (2003) DEN1 is a dual function protease capable of processing the C-terminus of Nedd8 deconjugating hyper-neddylated CUL1 [J]. J Biol Chem 278: 28882–28891.
[8] Walden H, Podgorski MS, et al. The structure of the APPBP1-UBA3-NEDD8-ATP complex reveals the basis for selective ubiquitin-like protein activation by an E1 [J]. Mol Cell. 12:1427–1437. 2003.
[9] Walden H, Podgorski M, Schulman BA. Insights into the ubiquitin transfer cascade from the structure of the activating enzyme for NEDD8 [J]. Letters to Nature. 2003;422:1-5.
[10] Huang DT, Hunt HW, et al. Basis for a ubiquitin-like protein thioester switch toggling E1–E2 affinity [J]. Nature. 2007;445:394-398.
[11] Walden H, Podgorski MS, et al. The structure of the APPBP1-UBA3-NEDD8-ATP complex reveals the basis for selective ubiquitin-like protein activation by an E1. Mol Cell [J]. 2003;12:1427–37.
[12] K. Chairatvit, C. Ngamkitidechakul. Control of cell proliferation via elevated NEDD8 conjugation in oral squamous cell carcinoma [J]. Mol. Cell. Biochem., 306 (2007), pp. 163-169.
CUSABIO team. NEDD8, An Important Target in Adipogenesis and Cancer Treatment. https://www.cusabio.com/c-19797.html



Comments
Leave a Comment