Tumor protein p53, encoded by the TP53 gene, is a tumor suppressor protein that maintains genomic stability and prevents cancer development. Often called the "guardian of the genome," p53 responds to cellular stresses including DNA damage, oncogene activation, and hypoxia. It triggers cell cycle arrest, DNA repair mechanisms, or apoptosis. TP53 mutations rank among the most frequent genetic alterations in human cancers, occurring in approximately 50% of all malignancies. This makes p53 one of the most studied proteins in cancer research.
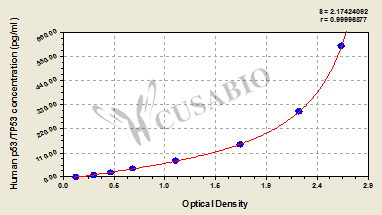
The Human p53/tumor protein,p53/TP53 ELISA Kit (CSB-E08334h) provides quantitative measurement of tumor protein p53 in human samples using a sandwich assay principle. This kit works with multiple sample types including serum, cell culture supernates, urine, cerebrospinal fluid, tissue homogenates, and cell lysates. The assay offers a detection range of 9.38 pg/mL to 600 pg/mL with a sensitivity of 2.34 pg/mL, requiring 50-100 μL sample volume and 1-5 hours assay time with detection at 450 nm wavelength.
Application Examples
Note: The following application examples are drawn from a selection of publications citing this product. For additional applications, please refer to the full list of references in the "Citations" section.
This ELISA kit has been used in cardiovascular research to quantify tumor protein p53 levels in cell culture studies examining cardiac damage mechanisms. The kit allows researchers to measure p53 secretion from human cardiomyocytes under various experimental conditions, particularly in models studying cellular stress responses and damage pathways.
• Cardiac damage research: Quantification of p53 levels in human cardiomyocyte culture supernatants to measure cellular responses to ischemia-reperfusion injury and other cardiac stress conditions
• miRNA functional studies: Analysis of p53 secretion changes following microRNA transfection experiments to understand regulatory mechanisms in cardiac cells
• Biomarker development: Integration with multi-analyte detection platforms for comprehensive biomarker profiling in research applications
• Cell culture studies: Measurement of p53 protein levels in controlled laboratory settings to study cellular damage responses and survival pathways