Histone Phosphorylation
In the 1970s, researchers began to detect phosphorylated histones in various biological samples. Experiments using radiolabeling techniques revealed that histones, particularly histone H1, could be phosphorylated. During the 1980s, studies started to suggest a link between histone phosphorylation and chromatin structure and function. One of the major breakthroughs in understanding histone phosphorylation came in the 1990s when researchers discovered its role in cell cycle regulation. In the 21st century, research into histone phosphorylation expanded significantly.
1. What Is Histone Phosphorylation?
Histone phosphorylation was first described in 1967 [1]. Histone phosphorylases deposit phosphate groups onto serine (S), threonine (T), or tyrosine (Y) residues on histones. Most phosphorylation occurs on H3S10 and H3S28, as well as H3T3, H3T6, and H3T11. Histone phosphorylation can have different effects on gene expression depending on the specific context and the location of the phosphorylation site.
2. Three Essential Components Involved in Histone Phosphorylation
Each of the four core histones is susceptible to phosphorylation by different protein kinases, and phosphorylation can be reversed by phosphatases, a process known as dephosphorylation. The recognition and binding of phosphorylated histones as well as ultimate translation of the signal into different biological outputs are accomplished by histone phosphorylation readers.
2.1 Writers
Protein kinases, also called writers, are enzymes that add phosphate groups to histones.
Table 1. Sites of histone phosphorylation, their known kinases and functions [2,16]
| Histone |
Phosphorylation Site |
Organism |
Kinase |
Biological Significance |
| H2A |
S1 |
Mammals |
MSK |
Mitosis, inhibition of transcription |
| T119 |
Mammals |
NHK-1,
AURKB |
Regulation of chromatin structure and function during mitosis |
| S121 |
Fission |
Bub1 |
Maintaining chromosomal homeostasis by recruiting shugoshin at sister kinetochore |
| S129 |
Yeast |
Mec1,
Tel1 |
DNA double-strand break repair |
| S139 (H2AX) |
Mammals |
ATR,
ATM,
DNA-PKcs,
RSK2,
MSK1 |
DNA repair, descreased EGF-mediated cellular transformation, apoptosis |
| Y142 (H2AX) |
Mammals |
WSTF |
Decision between cell survive and apoptosis |
| H2B |
S10 |
Yeast |
Ste20 |
Apoptosis |
| S14 |
Various vertebrates |
MstI |
Apoptosis |
| S32 |
Mammals |
PKC |
EGF signaling |
| S33 |
Drosophila |
CTK-TAF1 |
Transcriptional regulation |
| S36 |
Mammals |
AMPK |
Direct transcriptional and chromatin regulatory pathways leading to cellular response to stress |
| H3 |
T3 |
Mammals |
Haspin |
Mitosis |
| T6 |
Mammals |
PKCβ |
Androgen-dependent H3T6 phosphorylation prevents LSD1-mediated H3K4 demethylation, maintaining hormone-dependent gene activation |
| S10 |
Yeast, mammals |
Snf1,
IpL1
(yeast), AURKB (mammals), MSK1, MSK2, IKKα,
PKB/Akt, Rsk2, PIM1 |
Transcription, Chromatin condensation, UVB response |
| T11 |
Mammals |
Chk1,
PRK1, Dlk/Zip kinase |
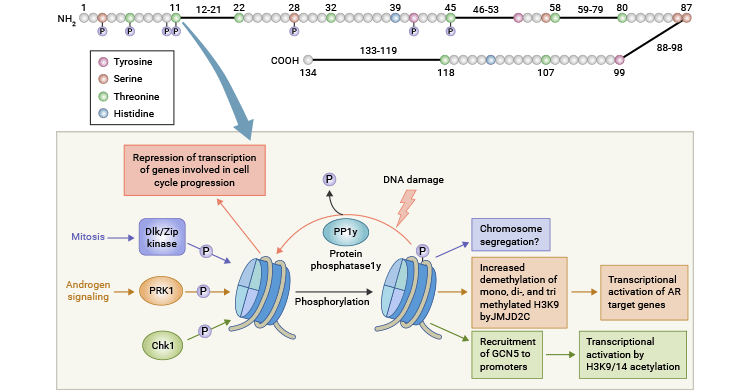
Meiosis, Transcription, DNA damage response |
|
|
| S28 |
Mammals |
AURKB, ERK1/ERK2, p38 |
Meiosis |
| MLTK-α, JNK1/2, MSK1 |
Mitosis, Transcription |
| Y41 |
Mammals |
JAK2 |
Transcription |
| T45 |
Mammals, budding yeast |
PK-Cδ,
Cdc7-Dbf4 |
Apoptosis, role in DNA-damaged cells when the DNA is nicked, replication of DNA |
| H4 |
S1 |
Yeast, mammals |
CKII |
DNA repair, Transcription |
| Sps1 |
Meiosis and chromatin assembly, Transcription |
| S47 |
Yeast, mammals |
PAK2 |
(H3.3-H4) Deposition |
| H18/75 |
Mammals |
- |
Facilitation of DNA replication by destabilizing histone octamer |
| H1 |
S/T |
|
CDK2 |
Mitosis, Transcription |
2.2 Erasers
Histone phosphatases, also known as erasers, remove phosphate groups from phosphorylated histones. There are eight distinct Ser/Thr-specific protein phosphatases (PPs) in mammals, including PP1, PP2A, PP2B, PP2C, PP4, PP5, PP6, and PP7 [3]. Among these, PP1, PP2A, and PP4 have all been identified as histone phosphatases.
| Histone phosphatases |
Recognized Dephosphorylation |
Organism |
| PP1 |
H1 [4] |
cell-cycle-dependent |
| PP1 homologue |
H3S10 [5] |
yeast and worms, during mitotic |
| PP2A |
H3S10ph [6] |
Drosophila |
| γ-H2AX [7] |
mammals and yeasts |
| PP4 |
γ-H2AX [8] |
mammals and yeasts |
2.3 Readers
Histone phosphorylation readers are phospho-histone binding proteins that recognize and bind to phosphorylated histones, thus interpreting the signals into different biological outputs. These proteins harbor phospho-binding domains, such as 14-3-3 and BRCT domains.
| Phospho-binding domain |
Reader |
Recognition Site |
Function |
| BRCT domain |
MDC1 |
H2A.X-S139 |
in response to DNA damage [9] |
| BIR domai |
survivin |
H3T3ph |
recruits chromosomal passenger complex to centromeres during mitosis [10-12] |
| 14-3-3 |
14-3-3ε,14-3-3ζ |
H3S10ph and H3S28ph |
linked to transcriptional regulation |
3. The Mechanism of Histone Phosphorylation
Understanding histone phosphorylation requires a grasp of its underlying mechanics:
3.1 Kinase Activation
Histone phosphorylation begins with the activation of kinases. These kinases can be activated in response to various signals, including cellular stress, growth factors, and DNA damage.
3.2 Phosphate Addition
Once activated, kinases target specific amino acids on histone proteins, mainly on serine, threonine, and tyrosine residues. Phosphate groups are added to these amino acids, altering the charge and structure of the histone.
3.3 Functional Outcomes
Histone phosphorylation affects chromatin structure and gene expression through several mechanisms. It can either create binding sites for other proteins or directly alter the interactions between histones and DNA. These changes ultimately dictate whether genes in the vicinity will be activated or silenced.
4. Function of Histone Phosphorylation
Histone phosphorylation and dephosphorylation, orchestrated by kinases and phosphatases, have gained recognition as a crucial mechanism for regulating a variety of chromatin-related processes, including transcription transcription, DNA replication, DNA damage repair, chromosome segregation, and apoptotic responses.
For example, H2A.X S139 and H4S1 are associated with DNA damage response. H3S10, H3S28, H2BS32, H3T6, H3T11, H3Y41, H2BS36, H2BY37, H4S1 and H4S47 are linked to transcription. H3S10 and H3S28 are found in mitosis. H4S1 and H2BS10 are observed in meiosis. H2BS10, H2AXS139, and H3T45 are involved in apoptosis.
Histone phosphorylation also plays an important role in the regulation of the cell cycle. In the G1 phase, histone phosphorylation promotes cell growth and DNA replication. During the S phase, histone phosphorylation helps maintain the stability of the replication fork. During G2 and M phases, histone phosphorylation is critical for chromosome separation and mitotic processes.
Phosphorylation of the variant histone H2A(X) at serine 139, known as γH2A(X), is a critical event in the DNA damage response (DDR). This modification takes place throughout all cell cycle phases and in response to various DDR pathways, including nonhomologous end joining (NHEJ), homologous recombination (HR), and DNA repair associated with replication.
Figure 1. Histone H2AX phosphoryaltion sites and important events occurring at S139 in mammalian cells [16]
5. Crosstalk between Histone Phosphorylation and Other Histone Modifications
Histone phosphorylation, similar to other histone modifications, does not act independently but rather participates in a complex interplay with other post-translational modifications, including methylation and acetylation.
H3S10 phosphorylation is physically and functionally coupled to acetylation of the nearly K14 residue, and acetylation of H3K14 enhances the interaction between H3S10ph and 14-3-3. Zippo A et al. found that H3S10ph can interact with the 14-3-3 protein and enabled the recruitment of the histone acetyltransferase MOF, which acetylates H3K16. These sequential events recruited Brd4 and P-TEFb, ultimately facilitating local transcription [13].
H3S28ph, in conjunction with H3K27ac, contributes to the activation of transcription. It achieves this by displacing Polycomb repressive complexes from chromatin, initiating demethylation and subsequent acetylation of the nearby H3K27 residue. This series of events ultimately results in the activation of transcription.
Phosphorylation of H3T6 by PKCβI inhibits the demethylation of H3K4 by LSD1 and JARID1B, while PRK1-mediated phosphorylation of H3T11 promotes the demethylation of H3K9 by JMJD2C [14, 15].
A study showed that H3T11 phosphorylation increased its interaction with the KAT Gcn5 at promoters of Gcn5-dependent genes, such as the cell-cycle regulators cyclin B and cdk1, resulting in heightened acetylation of H3K9 and K14, ultimately driving the activation of transcription.
The H4 acetyltransferase MOF is recruited to phosphorylated regions of H3S10 through its interaction with 14-3-3, resulting in crosstalk between H3S10ph and H4K16ac during the activation of immediate-early (IE) genes [13].
6. Histone Phosphorylation and Diseases
Histone phosphorylation is closely related to the occurrence and development of many diseases, including tumors, cardiovascular diseases, and neurological diseases. In tumors, abnormal histone phosphorylation leads to dysregulation of gene expression and promotes abnormal cell proliferation and metastasis. Abnormal phosphorylation of several histone proteins, as well as mutations in genes responsible for regulating histone phosphorylation, have been documented in numerous cancer types. In cardiovascular disease, an imbalance of histone phosphorylation can cause cardiomyocyte apoptosis and myocardial infarction. In neurological diseases, dysregulation of histone phosphorylation can lead to neuronal death and cognitive decline.
References
[1] Stevely WS, and Stocken LA (1966) Phosphorylation of rat-thymus histone [J]. Biochem. J 100, 20C-21C.
[2] Lau, P.N.I., and Cheung, P. (2009). Histone Phosphorylation: Chromatin Modifications that Link Cell Signaling Pathways to Nuclear Function Regulation [J]. In Handbook of Cell Signaling 2nd edition, R. A. Bradshaw and E. A. Dennis, eds. (Oxford: Academic Press), pp. 2399–2408.
[3] Moorhead GB, Trinkle-Mulcahy L, Ulke-Lemee A (2007). Emerging roles of nuclear protein phosphatases [J]. Nat Rev Mol Cell Biol 8: 234–244.
[4] Paulson JR, Patzlaff JS, Vallis AJ (1996). Evidence that the endogenous histone H1 phosphatase in HeLa mitotic chromosomes is protein phosphatase 1, not protein phosphatase 2A [J]. J Cell Sci 109: 1437–1447.
[5] Hsu JY et al. (2000). Mitotic phosphorylation of histone H3 is governed by Ipl1/aurora kinase and Glc7/PP1 phosphatase in budding yeast and nematodes [J]. Cell 102: 279–291.
[6] Nowak SJ, Pai CY, Corces VG (2003). Protein phosphatase 2A activity affects histone H3 phosphorylation and transcription in Drosophila melanogaster [J]. Mol Cell Biol 23: 6129–6138.
[7] Chowdhury D, Keogh MC, et al. (2005) γ-H2AX dephosphorylation by protein phosphatase 2A facilitates DNA double-strand break repair [J]. Mol Cell 20: 801–809.
[8] Chowdhury D, Xu X, et al. (2008) A PP4-phosphatase complex dephosphorylates γ-H2AX generated during DNA replication [J]. Mol Cell 31: 33–46.
[9] Stucki, M., Clapperton, et al. (2005). MDC1 directly binds phosphorylated histone H2AX to regulate cellular responses to DNA double-strand breaks [J]. Cell 123, 1213–26.
[10] Kelly, A.E., Ghenoiu, C., et al. (2010). Survivin Reads Phosphorylated Histone H3 Threonine 3 to Activate the Mitotic Kinase Aurora B [J]. Science 330, 235–239.
[11] Wang, F., Dai, J., et al. (2010). Histone H3 Thr-3 Phosphorylation by Haspin Positions Aurora B at Centromeres in Mitosis [J]. Science 330, 231–235.
[12] Yamagishi, Y., Honda, T., Tanno, Y., and Watanabe, Y. (2010). Two Histone Marks Establish the Inner Centromere and Chromosome Bi-Orientation [J]. Science 330, 239– 243.
[13] Zippo A, Serafini R, Rocchigiani M et al. (2009) Histone crosstalk between H3S10ph and H4K16ac generates a histone code that mediates transcription elongation [J]. Cell 138:1122–1136.
[14] Metzger, E., Yin, N., et al. (2008). Phosphorylation of histone H3 at threonine 11 establishes a novel chromatin mark for transcriptional regulation [J]. Nature Cell Biology 10, 53–60.
[15] Metzger, E., Imhof, A., et al. (2010). Phosphorylation of histone H3T6 by PKC|[bgr]|I controls demethylation at histone H3K4 [J]. Nature 464, 792.
[16] Banerjee T, Chakravarti D. A peek into the complex realm of histone phosphorylation. Molecular and Cellular Biology [J]. 2011 Dec;31(24):4858-4873.