Histone Ubiquitylation
The first hints of histone ubiquitylation came in the 1975 when researchers detected ubiquitin attached to histone H2A in the nucleosomes of calf thymus chromatin [1]. In the early 2000s, breakthrough research led to the identification of specific enzymes responsible for histone ubiquitylation. Later, researchers began to unravel the roles of histone ubiquitylation.
1. What Is Histone Ubiquitylation?
The process in which a 76-amino acid ubiquitin (Ub) moiety is incorporated into lysine residues of histone is known as histone ubiquitylation. Histone ubiquitylation includes histone mono-ubiquitylation and histone poly-ubiquitylation, depending on the added number of ubiquitin (Ub) moieties. Histone, particularly H2A and H2B, mainly occurs mono-ubiquitylation, which is a reversible process that alters the histone mass and influences nucleosomal dynamics. However, histone poly-ubiquitylation generates an irreversible signal for proteasomal-mediated degradation.
In most higher eukaryotes, 5%-15% of the total H2A proteins are ubiquitinated, while the H2B ubiquitylation level accounts for 1%-1.5% [1,2]. About 10% of H2B is ubiquitinated in budding yeast Saccharomyces cerevisiae, which contains no detectable levels of ubiquitinated H2A [2,3].
Histone H2A is mainly ubiquitinated at lysine 119 (H2AK119ub1), and the major H2B ubiquitylation sites are at lysine 123 (H2BK123ub1) in yeast or at lysine 120 (H2BK120ub1) in mammals [1,2]. Chromatin immunoprecipitation (ChIP) assays revealed that monoubiquitinated H2A is concentrated in the satellite regions of the genome, whereas H2Bub is localized across the body of active genes.
Beyond H2A and H2B, ubiquitin modification has been documented on core histones H3, H4, and linker histone H1. For instance, following UV irradiation, the CUL4-DDB-RBX1 ubiquitin ligase complex was found to polyubiquitinate H3 and H4 in vivo [4].
2. Mechanism of Histone Ubiquitylation
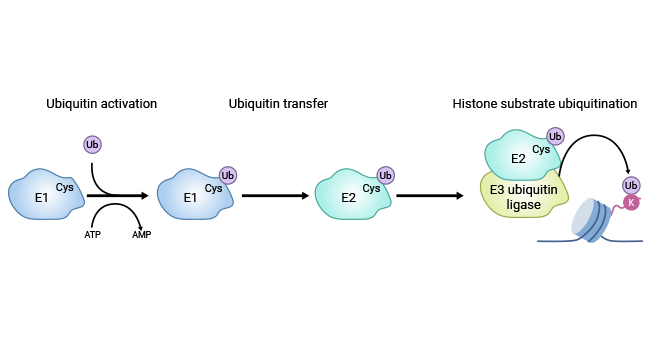
Ubiquitin is first attached to the E1 enzyme and activated in an ATP-dependent manner by forming a thiol-ester bond between its C-terminal glycine and the active cysteine residue of E1. Next, activated ubiquitin is transferred to the E2 enzyme through a thiol-ester linkage with ubiquitin. Finally, E2 interacts with an E3 ubiquitin ligase that covalently attaches the ubiquitin to a target lysine residue in the histone. The E3 ubiquitin ligase functions to position the E2-ubiquitin complex in proximity to the target lysine.
Figure 1. The process of histone mono-ubiquitylation.
3. Histone Ubiquitylation Enzymes
Histone ubiquitylation occurs through a series of orchestrated steps involving E1 activating, E2 conjugating, and E3 ligase enzymes. These steps culminate in the covalent bonding of ubiquitin (Ub) to a lysine (Lys) residue on histone proteins.
Histone ubiquitylation is a reversible process. Deubiquitinases (DUBs) remove ubiquitin from lysine residue on ubiquitinated histones.
| Enzyme |
Species |
Histone specificity |
Enzymatic activity |
Role in transcription |
| RING1A/RING1B/BMI1 |
Human |
H2A |
E3 |
Repression |
| 2A-HUB |
Human |
H2A |
E3 |
Repression |
| BRCA1/BARD1 |
Human |
H2A |
E3 |
Repression |
| UbcH5c |
Human |
H2A |
E2 |
N/A |
| Bre1 |
Yeast |
H2B |
E3 |
Activation |
| Rad6 |
Yeast |
H2B |
E2 |
Activation |
| RNF20/RNF40 |
Human |
H2B |
E3 |
Activation |
| RAD6A/RAD6B |
Human |
H2B |
E2 |
Activation |
| UbcH6 |
Human |
H2B |
E2 |
Activation |
| USP16 |
Human |
H2A |
DUB |
Activation |
| USP21 |
Human |
H2A |
DUB |
Activation |
| 2A-DUB |
Human |
H2A |
DUB |
Activation |
| BAP1 |
Human/Drosophila |
H2A |
DUB |
Activation |
| Ubp8 |
Yeast |
H2B |
DUB |
Activation |
| Ubp10 |
Yeast |
H2B |
DUB |
Repression |
| Ubp7 |
Drosophila |
H2B |
DUB |
Repression |
| SCNY |
Drosophila |
H2B |
DUB |
Repression |
| UBP12/UBP46
|
Xenopus |
H2A H2B |
DUB |
Activation |
| USP3 |
Human |
H2A H2B |
DUB |
N/A |
| USP22 |
Human |
H2A H2B |
DUB |
Activation |
| BAP1 |
Human/Drosophila |
H2A |
DUB |
Activation |
| Ubp8 |
Yeast |
H2B |
DUB |
Activation |
Table 1. Functions of histone modifying enzymes for monoubiquitylation.
The information is cited from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3355875/
Different from other types of histone modification readers, there isn't a shared domain or structural motif among readers of ubiquitylated histones.
| Modification |
Readers |
| H2AK119ub1 |
PRC1,
PRC2,
RSF1,
DNMT3A,
ZRF1 |
| H2AK13/15ub1 |
53BP1,
RNF169,
RAD18,
BARD1 |
| H2AK127/129ub1 |
SMARCAD1,
USP48 |
| H2BK120ub1 |
Dot1L,
COMPASS, MLL complexes, FACT, SWI/SNF, Chd1 |
| H2BK18/23ub1 |
DNMT1 |
| H3K14ub1 |
Clr4/SUV39H1 |
| H3K23/36/37ub1 |
Gcn5 |
Table 2. Histone ubiquitylation sites and cognate reader proteins.
The table information is from: https://www.frontiersin.org/articles/10.3389/fcell.2022.968398/full
4. Function of Histone Ubiquitylation
Given that histones stand as the most plentiful ubiquitinated proteins, their ubiquitylation assumes vital roles in almost all the DNA-related processes, such as transcription initiation and elongation, maintenance of chromatin structure, DNA replication, and DNA repair. The diverse consequences primarily hinge on the site-specific recognition of histone ubiquitylation by factors related to chromatin.
Mono-ubiquitylation of histones can bring about either gene activation and repression, whereas polyubiquitylation labels them for proteolytic degradation.
H2A119ub1 regulates gene repression at tissue-specific loci, at the inactive female X chromosome, or at regions of DNA damage. H2AK119 ubiquitylation is also involved in linker histone H1 binding to the nucleosome [5], Polycomb silencing [6], and female mammals’ X chromosome inactivation [7].
H2Bub1 participates in transcriptional activation and elongation, DNA damage response (DDR), and stem cell plasticity. H2Bub1 is a prerequisite for the methylation of H3K4 and H3K79 in both budding yeast and human cells [8,9]. Studies have demonstrated that H2Bub1 directly enhances the activity of the DOT1L methyltransferase, leading to the promotion of H3K79 methylation in both yeast and human models.
Histone ubiquitylation is also involved in sperm formation and plays an important role in the removal of nucleosomes during spermatogenesis.
5. Crosstalk between Histone Ubiquitylation and Other Histone Modifications
The crosstalk between histone ubiquitylation and other histone modifications represents a sophisticated regulatory network that governs chromatin dynamics and gene expression.
Histone lysine methylation has been demonstrated to be cross-regulated by histone ubiquitylation or histone ubiquitylation modifying enzymes. Monoubiquitinated H2A is recognized by PRC2, causing H3K27 methylation [13]. This recruits SETDB1 for H3K9 trimethylation, anchoring chromatin compactors like HP1 [14,15]. Trimethylated H3K27 attracts more PRC1, initiating stable long-range signaling via H2A modification [16]. The monoubiquitylation of H2B triggers DOT1L to methylate histone H3 at K79, subsequently enhancing H3K4 methylation by MLL, which facilitates the recruitment of transcriptional activators [17].
Histone ubiquitylation also shows crosstalk with lysine acetylation in the DNA damage response. Knocking down lysine acetyltransferases KAT5 (TIP60) or PCAF led to a reduction in the acetylation of lysine 120 on H2B (H2BK120ac), along with an elevation in the ubiquitylation of H2BK120, in response to DNA damage [10]. EP300 acetylates the H3K9me2 demethylase JMJD1A on lysine 421, which prevents STUB1-mediated polyubiquitylation, consequently preserving JMJD1A levels [11]. H2B monoubiquitylation also interacts with histone H3 methylation, promoting gene transcription and telomeric silencing.
There are documented interrelationships between ubiquitylation and phosphorylation in chromatin control. EGFR triggers the attachment of lysine 48-linked polyubiquitin chains to H3K4 through RNF8-UBE2L6, leading to the degradation of H3 [12]. When threonine 11 on H3.3 (H3.3T11ph) was mutated to alanine, both phosphorylation and ubiquitylation were blocked. Histone phosphorylation and ubiquitylation collaborate synergistically during DNA damage signaling triggered by ionizing radiation (IR).
6. Histone Ubiquitylation and Diseases
Dysregulation of histone ubiquitylation or deubiquitylation has been regarded as a causative factor for various human diseases, including brain-associated disorders and cancer.
Mutations affecting various components involved in regulating H2Aub levels have been correlated with several syndromes and brain-related disorders. For example, mutations within the AUTS2 gene are connected to autism and intellectual disability. Deficiencies in PHC1 are associated with disorders in brain development, like autosomal recessive primary microcephaly. PHC1 mutations have been linked to compromised H2A ubiquitylation, disruptions in cell cycle progression, and impaired DNA repair mechanisms.
Loss of PRC1 is incompatible with life. Mutations that hinder PRC1's activity lead to microcephaly or learning disabilities. Abnormal modulation of PRC1 also causes tumors in multiple organs, including the brain, liver, colon, breast, lung, prostate, or lymphatic system.
Irregular mono-ubiquitylation of H2Bub1 is frequently associated with tumorigenesis. RNF20/40 is commonly disrupted in primary tumors like colorectal, breast, ovarian, prostate, and lung cancers. The deficiency of H2Bub1 often corresponds with unfavorable prognoses among cancer patients [18].
Amplification or excessive expression of RING and BMI1 results in elevated levels of H2Aub and tumor progression. Cancer mutations within BAP1's UCH, CTD, and NLS functional domains lead to H2Aub deubiquitylation loss and contribute to cancer development.
References
[1] Goldknopf I. L., Taylor C. W., et al. (1975). Isolation and characterization of protein A24, a “histone-like” non-histone chromosomal protein [J]. J. Biol. Chem. 250, 7182–7187.
[2] West M. H., Bonner W. M. (1980). Histone 2B can be modified by the attachment of ubiquitin [J]. Nucleic Acids Res. 8, 4671–4680.
[3] Robzyk K., Recht J., Osley M. A. (2000). Rad6-dependent ubiquitylation of histone H2B in yeast [J]. Science 287, 501–504.
[4] Wang H., Zhai L., et al. (2006). Histone H3 and H4 ubiquitylation by the CUL4-DDB-ROC1 ubiquitin ligase facilitates cellular response to DNA damage [J]. Mol. Cell 22, 383–394.
[5] Jason LJ, Finn RM, et al. Histone H2A ubiquitylation does not preclude histone H1 binding, but it facilitates its association with the nucleosome [J]. J. Biol. Chem. 2005;280:4975–4982.
[6] Wang H, Wang L, et al. Role of histone H2A ubiquitylation in Polycomb silencing [J]. Nature. 2004;431:873–878.
[7] Fang J, Chen T, et al. Ring1b-mediated H2A ubiquitylation associates with inactive X chromosomes and is involved in initiation of X inactivation [J]. J. Biol. Chem. 2004;279:52812–52815.
[8] Kim J, Guermah M, et al. RAD6-mediated transcription-coupled H2B ubiquitylation directly stimulates H3K4 methylation in human cells [J]. Cell. 2009;137:459–471.
[9] Nakanishi S, Lee JS, et al. Histone H2BK123 monoubiquitylation is the critical determinant for H3K4 and H3K79 trimethylation by COMPASS and Dot1 [J]. J. Cell Biol. 2009;186:371–377.
[10] Kim J.J. et al. Systematic bromodomain protein screens identify homologous recombination and R-loop suppression pathways involved in genome integrity [J]. Genes Dev. 2019; 33: 1751-1774.
[11] Xu S. et al. p300-mediated acetylation of histone demethylase JMJD1A prevents its degradation by ubiquitin ligase STUB1 and enhances its activity in prostate cancer [J]. Cancer Res. 2020; 80: 3074-3087.
[12] Xia Y. et al. RNF8 mediates histone H3 ubiquitylation and promotes glycolysis and tumorigenesis [J]. J. Exp. Med. 2017; 214: 1843-1855.
[13] Kalb R, Latwiel S, et al. (2014) Histone H2A monoubiquitylation promotes histone H3 methylation in Polycomb repression [J]. Nat Struct Mol Biol 21: 569–571.
[14] Zhao D, Zhang X, et al. (2016) The BAH domain of BAHD1 is a histone H3K27me3 reader [J]. Protein Cell 7: 222–226.
[15] Machida S, Takizawa Y, et al. (2018) Structural basis of heterochromatin formation by human HP1 [J]. Mol Cell 69: 385–397.e8.
[16] Tamburri S, Lavarone E, et al. (2020) Histone H2AK119 mono-ubiquitylation is essential for polycomb-mediated transcriptional repression [J]. Mol Cell 77: 840–856.e5.
[17] Wu L, Zee BM, et al. (2011) The RING finger protein MSL2 in the MOF complex is an E3 ubiquitin ligase for H2B K34 and is involved in crosstalk with H3 K4 and K79 methylation [J]. Mol Cell 43: 132–144.
[18] Sa Zhou, Yuqiao Cai, et al. Role of H2B mono-ubiquitylation in the initiation and progression of cancer [J]. Bulletin du Cancer, Volume 108, Issue 4, April 2021, Pages 385-398.