Mutations in Spike Protein of SARS-CoV-2
As mentioned on the article entitled “An Overview of 2019 Novel Coronavirus (SARS-CoV-2)”, SARS-CoV-2 is comprised of four structural proteins, including N (nucleocapsid), M (membrane), E (envelope) and S (spike). Among of them, S protein forms a knob like structure protruding outwards beyond the other structural proteins, and it plays a key role in SARS-CoV-2 entry. Accumulating evidences have showed mutations in structural proteins of virus play crucial role in viral virulence by determining generation of antibody escape variants and cellular tropism. Moreover, mutation is the basis of virus evolution, and can make the virus escape immune response of the host, and can be more conducive to the spread of the virus. Here, we focus on the mutations in Spike (S) protein of SARS-CoV-2.
What is The Spike Protein of SARS-CoV-2?
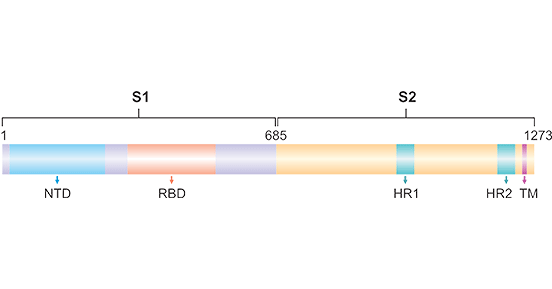
Spike protein is one of the four structural proteins of SARS-CoV-2, and comprises two functional subunits, S1 subunit and S2 subunit. S1 subunit is responsible for binding to the host cell receptor. S2 submit is in charge of fusion of the viral and cellular membranes. S1 contains the receptor binding domain (RBD) and mediates virus attachment to its ACE2 receptor. S2 contains the fusion peptide. As the Figure 1 shows, the amino acids sequence from1 to 13aa is signal peptide; S1 is from 13 to 685aa, and 319-541aa is RBD. The amino acids sequence from 686-1273aa is S2, and 770-788aa is the fusion peptide (reference: UniRule: UR000875864)
Figure 1. Schematic showing structural organization of SARS-CoV-2 spike protein
*This diagram is derived from the publication published by Arup Kumar Banerjee [1]
How did The SARS-CoV-2 Mutants Attract Attention of Scientists?
On March 15, 2020, a research team from the West China Hospital of Sichuan University published an unpeer-reviewed article entitled "International expansion of a novel SARS-CoV-2 mutant" on medRxiv, reporting a new SARS-CoV-2 mutations seem to be spreading around the world and deserve close attention [2]. Then, the team of Fang Li from University of Minnesota studied the spike protein (S protein) on the surface of SARS-CoV-2 and attracted more attention to the SARS-CoV-2. The results of this study were published in Nature. They found that some mutations occurred in the SARS-CoV-2 strain, forming a particularly tight "ridge" on the S protein by X-ray crystallography. This ridge is more compact than that in the SARS virus, which may be one of the reasons why SARS-CoV-2 is so contagious [3]. From then on, researches on S protein mutants have been gradually carried out, and has achieved considerable results.
The Hot Spots Mutations in Spike Protein of SARS-CoV-2
Several evidences have showed that many mutations have been identified in the S protein, and there are also portions of S protein that remained unchanged and might be conserved. The mutations are mainly concentrated in the S1 subunit, especially in the RBD, and the conserved domains are primarily focused on the S2 subunit.
On April 9, 2020, Junxian Ou et al. preprinted a study entitled “Emergence of RBD mutations from circulating SARS-CoV-2 strains with enhanced structural stability and higher human ACE2 receptor affinity of the spike protein” on the bioRxiv, which was not certified by peer review. They found that 32 RBD mutant strains clustering into 10 mutation types under high positive selection pressure by analyzing genomes of global SARS-CoV-2 strains. Among of the 32 RBD mutant strains, three RBD mutation types (N354D and D364Y, V367F, W436R) exhibited significantly lowered ΔG, suggesting a significantly increased affinity to human ACE2, and indicating these mutants may have acquired increased infectivity to humans [4]. In the only double amino acid mutant (N354D, D364Y), the N354D substitution decreased the affinity, although the D364Y provided an even higher affinity than the overall double mutations. This indicated that the D364Y is the major contributor of the enhanced affinity.
On 16 April 2020,the study of Arup Kumar Banerjee et al. from India have pointed out three major hot spots of mutations (G476S, V483A, D614G) in the S1 subunit of SARS-CoV-2 isolated from the North America. Among of them, at position 614, mutation D to G is clearly a very frequent mutation, and this type of mutation doesn’t fall in the RBD of spike protein. The other two types of mutation (G476S, V483A) fall in the RBD of spike protein. And V483A repeated more frequently followed by G476S. Of course, beside these three types of mutation in the S1, there also are several hot spots of mutations in the S1, such as A348T, D111N and G176V, etc.
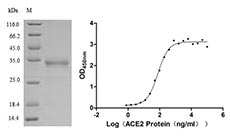
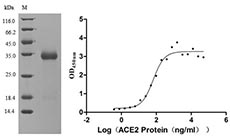
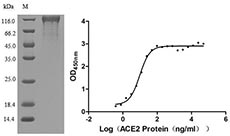
Now, CUSABIO have developed several SARS-CoV-2 Spike S1 mutation recombiant proteins, including RBD (V367F), RBD (W436R), RBD (G476S), RBD (V483A) and S1 (D614G). All of these proteins are validated in SDS-PAGE and function ELISA. The purity of them are more than 90%.
Spike Mutant Proteins of SARS-CoV-2
| Target Name |
Product Name |
Code |
Source |
Tag Info |
| S (RBD) |
Recombinant Human Novel Coronavirus Spike glycoprotein(S) (V367F), partialIn Stock |
CSB-MP3324GMY1(M1) |
Mammalian cell |
C-terminal 10xHis-tagged |
| S (RBD) |
Recombinant Human Novel Coronavirus Spike glycoprotein(S) (W436R), partialIn Stock |
CSB-MP3324GMY1(M2) |
Mammalian cell |
C-terminal 10xHis-tagged |
| S (RBD) |
Recombinant Human Novel Coronavirus Spike glycoprotein(S) (N354D,D364Y), partial |
CSB-MP3324GMY1(M3) |
Mammalian cell |
C-terminal 10xHis-tagged |
| S (RBD) |
Recombinant Human Novel Coronavirus Spike glycoprotein(S) (G476S), partialIn Stock |
CSB-MP3324GMY1(M4) |
Mammalian cell |
C-terminal 10xHis-tagged |
| S (RBD) |
Recombinant Human Novel Coronavirus Spike glycoprotein(S) (V483A), partialIn Stock |
CSB-MP3324GMY1(M5) |
Mammalian cell |
C-terminal 10xHis-tagged |
| S (RBD) |
Recombinant Human Novel Coronavirus Spike glycoprotein(S) (N501Y), partial (Active)In Stock |
CSB-MP3324GMY1(M6) |
Mammalian cell |
C-terminal 10xHis-tagged |
| S (RBD) |
Recombinant Human Novel Coronavirus Spike glycoprotein(S) (N501Y), partial (Active)In Stock |
CSB-MP3324GMY1(M6)k2 |
Mammalian cell |
C-terminal 6xHis-mFC-tagged |
| S (RBD) |
Recombinant Human Novel Coronavirus Spike glycoprotein(S) (K417N), partial |
CSB-MP3324GMY1(M7) |
Mammalian cell |
C-terminal 10xHis-tagged |
| S (RBD) |
Recombinant Human Novel Coronavirus Spike glycoprotein(S) (K417N), partial |
CSB-MP3324GMY1(M7)h8 |
Mammalian cell |
C-terminal mFC-tagged |
| S (RBD) |
Recombinant Human Novel Coronavirus Spike glycoprotein(S) (E484K), partial |
CSB-MP3324GMY1(M8) |
Mammalian cell |
C-terminal 10xHis-tagged |
| S (RBD) |
Recombinant Human Novel Coronavirus Spike glycoprotein(S) (E484K), partial (Active)In Stock |
CSB-MP3324GMY1(M8)h8 |
Mammalian cell |
C-terminal mFC-tagged |
| S (S1) |
Recombinant Human Novel Coronavirus Spike glycoprotein(S) (D614G), partialIn Stock |
CSB-MP3324GMY(M1) |
Mammalian cell |
C-terminal 10xHis-tagged |
Nucleoprotein Mutant Protein of SARS-CoV-2
Biotinylated Protein of SARS-CoV-2
References
[1] Arup Kumar Banerjee, Feroza Begum, et al. Mutation hot spots in Spike protein of COVID-19 [J]. Preprints. 2020.
[2] Minjin Wang, Mengjiao Li, et al. International Expansion of a Novel SARS-CoV-2 Mutant [J]. J Virol. 2020, 94(12):e00567-20.
[3] Jian Shang, Gang Ye, et al. Structural basis of receptor recognition by SARS-CoV-2 [J]. Nature. 2020, 581, 221–224.
[4] Junxian Ou, Zhonghua Zhou, et al. Emergence of RBD mutations from circulating SARS-CoV-2 strains with enhanced structural stability and higher human ACE2 receptor affinity of the spike protein [J]. bioRxiv. 2020.