Hepatitis Viruses
Hepatitis viruses, including A, B, C, D, and E, pose a global health threat by inducing hepatitis. Type A and E typically spread through contaminated food and water, while B, C, and D transmit via infected blood and bodily fluids. These viruses can lead to acute or chronic liver diseases, with Hepatitis B and C often progressing stealthily to chronic conditions, increasing the risk of cirrhosis and liver cancer. All these hepatitis viruses are RNA viruses except HBV. Vaccines are available for Hepatitis A and B, contributing to prevention efforts.
Table 1. Characteristics of hepatitis viruses
|
HAV |
HBV |
HCV |
HDV |
HEV |
| Family |
Picornaviridae |
Hepadnaviridae |
Flaviviridae |
Not applicable |
Hepeviridae |
| Genus |
Hepatovirus |
Orthohepadnavirus |
Hepacivirus |
Deltavirus |
Orthohepevirus |
| Genome |
Positive-sense single-stranded linear RNA |
Double-stranded DNA |
Positive-sense single-stranded linear RNA |
Negative-sense single-stranded circular RNA |
Positive-sense single-stranded linear RNA |
| Transmission |
Fecal–oral |
Exposure to infected blood or body fluid/sexual/perinatal |
Exposure to infected blood |
Exposure to infected blood/body fluids |
Feca–oral/zoonotic/blood transfusion |
| Clinical outcome of infection |
Self-limited |
Self-limited and chronic |
Self-limited and chronic |
Self-limited and chronic |
Self-limited |
The Table information is cited from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC8999150/
Hepatitis A virus (HAV)
HAV is a non-enveloped RNA virus with a 7.5 kb genome. Its genome encodes a single large polyprotein that is processed into four capsid proteins (VP1, VP2, VP3, VP4) and seven non-structural polypeptides (2A, 2B, 2C, 3A, 3B, 3C, 3D). Infectious HAV virions exist in two forms: naked, non-enveloped HAV (neHAV) virions released into the stool, and quasi-enveloped virions (eHAV) generated by hijacking control of the host.
HAV Life Cycle
HAV enters host hepatocytes by engaging with cell surface molecules, specifically sialic acid and ganglioside, followed by the uncoating of viral capsid and subsequent release of viral RNA from endosomes into the cytoplasm [1]. The viral genome is translated into a polyprotein under the regulation of an internal ribosome entrance site (IRES) and then cleaved into distinct structural and nonstructural proteins.
The genomic RNA is also used as a template to generate numerous new replicas of the RNA genome, which are packed into capsids to produce intracellular viral offspring. Newly synthesized HAV virions are released from cells in quasi-enveloped forms.
Hepatitis B Virus (HBV)
The HBV virion consists of an outer lipoprotein envelope composed of hepatitis B surface antigen (HBsAg), and a hepatitis B core antigen (HBcAg)-formed icosahedral nucleocapsid which encapsulates a partially double-stranded, relaxed circular DNA (rcDNA) genome of 3.2 kb.
The HBV genome consists of four partially overlapping and open reading frames (ORFs), C (core), P (polymerase), X (regulatory X protein), and S (surface), from which seven functional proteins are synthesized: HBcAg, HBeAg, polymerase (pol), HBx, and three types of HBsAg - the small (s), medium (M), and large (L) proteins.
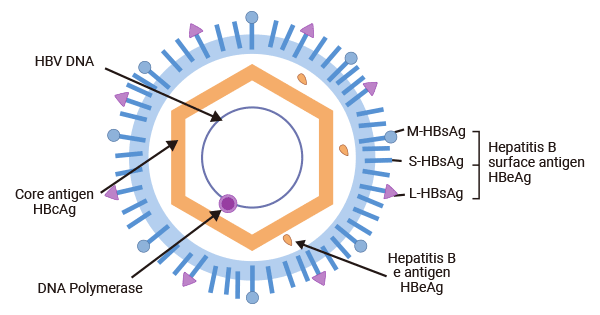
Figure 1. HBV structure
The picture is cited from Wikipedia
| HBV proteins |
Function |
Products |
| HBcAg |
Assemble intothe icosahedral viral nucleocapsid to enclose the HBV genome and DNA polymerase |
Recombinant HBV genotype A1 subtype adw Capsid protein (C) |
| Recombinant HBV genotype A2 Capsid protein (C) |
| Recombinant HBV genotype A3 Capsid protein (C) |
| Recombinant HBV genotype B1 Capsid protein (C) |
| Recombinant HBV genotype B2 Capsid protein (C) |
| Recombinant HBV genotype C Capsid protein (C) |
| Recombinant HBV genotype D Capsid protein (C) |
| Recombinant HBV genotype F2 Capsid protein (C) |
| Recombinant HBV genotype G Capsid protein (C) |
| Recombinant HBV genotype H subtype adw4 Capsid protein (C) |
| HBeAg |
Not necessary for viral replication or infection; an indicator of active viral replication and is linked to the establishment of chronic infection, hepatic inflammatory injury, and HCC development |
Recombinant HBV genotype A1 subtype adw External core antigen (C) |
| Recombinant HBV genotype A2 External core antigen (C) |
| Recombinant HBV genotype A3 External core antigen (C) |
| Recombinant HBV genotype B1 Truncated HBeAg protein (C) |
| Recombinant HBV genotype B2 Truncated HBeAg protein (C) |
| Recombinant HBV genotype C External core antigen (C) |
| Recombinant Hepatitis B virus genotype D Truncated HBeAg protein (C) |
| Recombinant HBV genotype F2 subtype adw4q External core antigen (C) |
| Recombinant HBV genotype H subtype adw4 External core antigen (C) |
| DNA Polymerase |
Initiate viral replication by reverse transcription |
Recombinant HBV genotype A1 subtype adw2 Protein P |
| Recombinant HBV genotype A2 Protein P |
| Recombinant HBV genotype A3 Protein P |
| Recombinant HBV genotype B1 Protein P |
| Recombinant HBV genotype B2 Protein P |
| Recombinant HBV genotype C Protein P |
| Recombinant HBV genotype D Protein P |
| Recombinant HBV genotype E Protein P |
| Recombinant HBV genotype F1 Protein P |
| Recombinant HBV genotype F2 Protein P |
| Recombinant HBV genotype G Protein P |
| Recombinant HBV genotype H Protein P |
| HBx |
Localized in different subcellular compartments, conferring distinct functions; involved in various cellular events, including transactivator of the viral and cellular promoters, epigenetic modifications, ubiquitination, autophagy and non-coding RNA regulation, to promote hepatocellular carcinogenesis, stabilization and activation of cccDNA |
Recombinant HBV genotype A1 subtype adw2 Protein X |
| Recombinant HBV genotype A2 Protein X |
| Recombinant HBV genotype A3 Protein X |
| Recombinant HBV genotype B1 Protein X |
| Recombinant HBV genotype B2 Protein X |
| Recombinant HBV genotype C Protein X |
| Recombinant HBV genotype D Protein X |
| Recombinant HBV genotype E Protein X |
| Recombinant HBV genotype F1 subtype adw4 Protein X |
| Recombinant HBV genotype F2 Protein X |
| Recombinant HBV genotype G Protein X |
| Recombinant HBV genotype H Protein X |
| HBsAg |
Include L-HBsAg, M-HBsAg, and S-HBsAg; make up the viral envelope |
Recombinant HBV genotype A1 subtype adw2 Large envelope protein (S) |
| Recombinant HBV genotype A2 subtype adw Large envelope protein (S) |
| Recombinant HBV genotype A3 Large envelope protein (S) |
| Recombinant HBV genotype B1 Large envelope protein (S) |
| Recombinant HBV genotype B2 Large envelope protein (S) |
| Recombinant HBV genotype C Large envelope protein (S) |
| Recombinant HBV genotype D Large envelope protein (S) |
| Recombinant HBV genotype E Large envelope protein (S) |
| Recombinant HBV genotype F1 Large envelope protein (S) |
| Recombinant HBV genotype F2 Large envelope protein (S) |
| Recombinant HBV genotype G Large envelope protein (S) |
| Recombinant HBV genotype H Large envelope protein (S) |
HBV Life Cycle
HBV enters into the host hepatocytes in an endocytosis-dependent manner. HBV initiates by binding to heparan sulfate proteoglycans (HSPGs) such as glypican 5. And then, it engages with the sodium taurocholate cotransporting peptide (NTCP/SLC10A1) receptor on the surface of hepatocytes to elicit viral internalization [2-4].
Studies have demonstrated that EGFR triggers HBV’s internalization into host cells by interacting directly with NTCP [5,6]. Following endocytosis, the viral nucleocapsid is released into the hepatic cytoplasm and then transported to the nucleus.
In the nucleus, the viral genomic rcDNA is converted to cccDNA (covalently closed circular DNA) to amplify a viral RNA intermediate, which is then reverse-transcribed back to viral DNA [7]. Simultaneously, the viral DNA integrates into the host genome, which is the cause that hepatitis B is incurable.
The cccDNA minichromosome remains in the nucleus, serving as a template for viral RNA transcription. It encodes seven essential viral proteins necessary for replication: three different sizes of HBsAg, HBcAg, HBeAg, HBx, and DNA polymerase [8].
In the cytoplasm, HBcAg proteins self-assemble into an icosahedral nucleocapsid, encapsulating viral polymerase and pgRNA. Following reverse transcription within the nucleocapsid, viral capsid that contains the HBV genome binds to HBsAg proteins in the endoplasmic reticulum for encapsulation. Ultimately, mature HBV virions are released from hepatocytes through multivesicular bodies (MVBs).
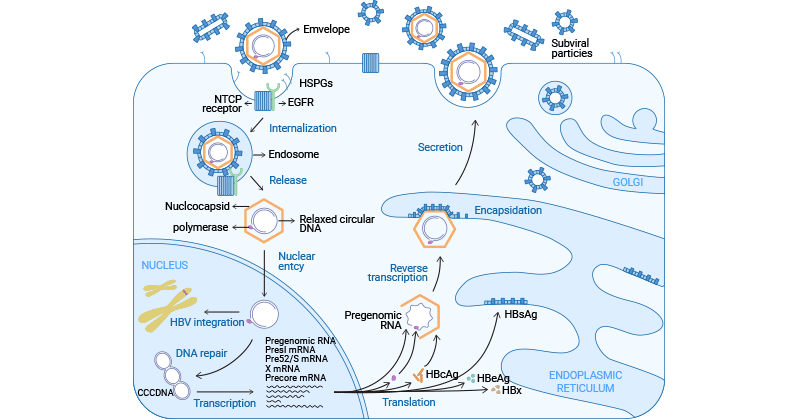
Figure 2. HBV life cycle
The picture is cited from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC8387624/
Hepatitis C Virus (HCV)
HCV has an RNA genome of about 9.6 kb in length. The viral genome contains a single open reading frame (ORF) that encodes a single polyprotein precursor, which is subsequently processed into three structural proteins and seven non-structural proteins.
HCV Life Cycle
HCV enters the cell in a pH and clathrin-dependent endocytosis. HCV first attaches to the host cells by binding to HSPG and LDL-R. After docking, the HCV particle binds to CD81 and scavenger receptor class B type I (SR-BI) on the surface of the hepatocyte through E2 [9,10] and then interacts with claudin-1 and occludin, which facilitates its entry into the cell with the endosome [11].
Following endosome-mediated internalization, endosomal acidification promotes low pH-dependent HCV envelope fusion with the endosomal membrane, resulting in the release of viral genomic RNA into the cytosol [12]. The RNA is used as the template both for replication and polyprotein translation. The RNA translation takes place in the ER and is initiated by binding the 5'UTR IRES to the ribosome, leading to the formation of the polyprotein precursor.
The polyprotein is co- and post-translationally cleaved by the host or viral proteases to yield the structural proteins (core, E1, and E2) and the nonstructural proteins (p7, NS2, NS3, NS4A, NS4B, NS5A, and NS5B proteins). HCV RNA replication occurs within the membranous web (MW) in the ER and is catalyzed by the NS5B RNA-dependent RNA polymerase. The newly generated viral RNA is transported to the site of core assembly and then is encapsulated to form the nucleocapsid.
The viral nucleocapsids subsequently interact with the viral E1/E2 proteins (envelopment) and bud into the ER lumen and then are translocated to the Golgi. The viral virions finish maturation in the Golgi and are transported to the plasma membrane through very-low-density lipoprotein (VLDL) pathway and released from host hepatocytes.
Hepatitis D Virus (HDV)
Rizzetto et al. initially discovered HDV in HBV-infected individuals with severe hepatitis in 1977 [13]. HDV is a hybrid virus as it uses Hepatitis B surface antigen (HBsAg) as its envelope protein. This characteristic allows HDV to infect only patients that concomitantly carry HBV.
HDV particle is composed of an outer lipoprotein envelope made of the surface antigen of the HBV (HBsAg) and an inner ribonucleoprotein structure in which the HDV genome resides. The HDV genome is about 1.7 kb and encodes only for the hepatitis delta antigen (HDAg) with two forms: a 27 kDa large-HDAg (delta-Ag-L), and a small-HDAg of 24 kDa (delta-Ag-S) [14].
HDV Life Cycle
HDV infection is dependent on the presence of the HBV as it requires HBV's envelope proteins for its entry, assembly, and transmission. HDV virions initially attach to HSPGs on the hepatocyte surface and subsequently enter host cells through NTCP receptor [15]. After viral envelope fusion with the endosomal membrane, HDV ribonucleoprotein (RNP) is released from the endosome and then transported to the cellular nucleus.
In the nucleolus, HDV RNA undergoes replication through a double rolling circle amplification mechanism, generating both antigenomic RNA and additional genomic RNA. After export to the cytoplasm, the mRNA is translated at the ER to generate HDAg proteins. HDAg proteins then re-enter the nucleus, where the S-HDAg enhances genome replication. S-HDAg and L-HDAg associate with new synthesized genomic RNA to form new RNPs.
The resulting RNPs are released to the cytoplasm, where L-HDAg aids in linkage to HBsAg in the ER to assemble new viral particles. The newly synthesized viral particles are subsequently released from the hepatocytes through the Golgi to infect adjacent cells.
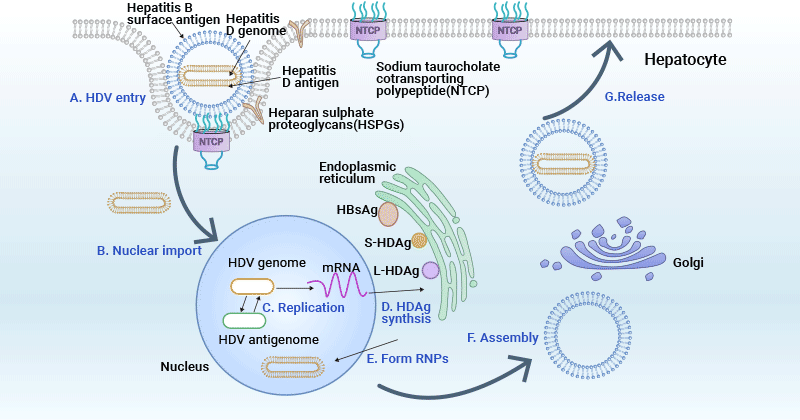
Figure 3. HDV life cycle
This picture is cited from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC8557527/
Hepatitis E Virus (HEV)
Similar to HAV, HEV virions exist in two forms in the infected host, non-enveloped (neHEV) spherical particles in feces and quasi-enveloped (eHEV) particles in circulating blood and supernatant of infected cell cultures.
The HEV genomic RNA is about 7.2 kb in length and has three open reading frames (ORFs): ORF1 encodes a nonstructural polyprotein for virus replication and transcription, ORF2 encodes the capsid protein that contains the S, M, and P domains, and ORF3, which partially overlaps ORF2, encodes a multifunctional phosphoprotein involved in virion morphogenesis and pathogenesis.
Recently, a novel ORF4 has been discovered only in genotype 1 HEV, and endoplasmic reticulum (ER) stress boosts viral replication by promoting the translation of the novel ORF4 in genotype 1 HEV [16].
HEV Life Cycle
HEV infection begins when an individual consumes feces-contaminated water or food, introducing the virus into the digestive system. HEV subsequently enters the bloodstream through viremia and reaches the liver. Non-enveloped HEV particles bind to HSPGs on the surface of hepatocytes and enter host cells via a specific cellular receptor. Quasi-enveloped HEV virions enter liver cells through dynamin-dependent, clathrin-mediated endocytosis, involving Rab5 and Rab7.
After entry into host hepatocytes, HEV particles undergo uncoating of capsid protein and then release its genomic RNA to the cytoplasm. The viral genomic RNA serves directly as mRNA for the translation of ORF1 polyprotein. It also synthesizes a complementary negative-sense RNA, which acts as a template for the transcription of genomic and subgenomic mRNA.
The subgenomic mRNA is then translated to ORF2 capsid protein and ORF3 multifunctional phosphoprotein. ORF2 capsid protein forms virus-like particles (VLPs) and packages newly synthesized positive-sense genomic RNA to produce progeny HEV virions.
ORF3 modulates the host environment by interacting with cellular proteins, facilitating viral replication and release. Notably, ORF3 associates with TSG101 in the ESCRT pathway, aiding the budding of nascent virions into multivesicular bodies. These multivesicular bodies fuse with the plasma membrane, releasing viral virions from infected hepatocytes, either into the bloodstream as eHEV or in the bile duct as neHEV.
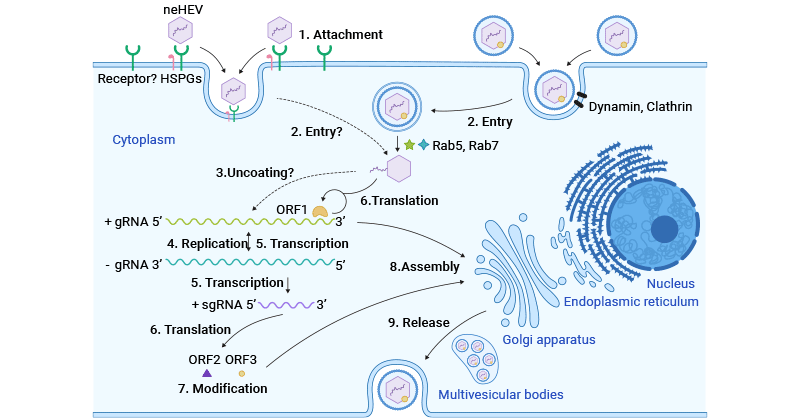
Figure 3. HEV life cycle
This picture is cited from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC8079827/
The global situation of hepatitis viruses remains a significant public health concern. Hepatitis B and C, in particular, pose ongoing challenges due to their ability to cause chronic infections leading to severe liver diseases. The development of effective antiviral therapies and vaccination programs has made progress in preventing and managing hepatitis infections. However, barriers to access, especially in low-income regions, persist.
References:
[1] Gholizadeh O, Akbarzadeh S, et al. Hepatitis A: Viral Structure, Classification, Life Cycle, Clinical Symptoms, Diagnosis Error, and Vaccination [J]. Can J Infect Dis Med Microbiol. 2023 Jan 4;2023:4263309.
[2] A. Schulze, P. Gripon, and S. Urban. Hepatitis B virus infection initiates with a large surface protein-dependent binding to heparan sulfate proteoglycans [J]. Hepatology, 46 (2007), pp. 1759-1768.
[3] E.R. Verrier, C.C. Colpitts, et al. A targeted functional RNA interference screen uncovers glypican 5 as an entry factor for hepatitis B and D viruses [J]. Hepatology, 63 (2016), pp. 35-48.
[4] H. Yan, G. Zhong, eet al. Sodium taurocholate cotransporting polypeptide is a functional receptor for human hepatitis B and D virus [J]. Elife, 1 (2012), Article e00049.
[5] M. Iwamoto, W. Saso, et al. Epidermal growth factor receptor is a host-entry cofactor triggering hepatitis B virus internalization [J]. Proc. Natl. Acad. Sci. U. S. A., 116 (2019), pp. 8487-8492.
[6] Seeger C, Mason WS. Molecular Biology of Hepatitis B Virus Infection [J]. Virology (2015) 479-480:672–86.
[7] Schreiner S, Nassal M. A Role for the Host DNA Damage Response in Hepatitis B Virus cccDNA Formation-And Beyond [J]? Viruses (2017) 9(5):125.
[8] Tong S, Revill P. Overview of Hepatitis B Viral Replication and Genetic Variability [J]. J Hepatol (2016) 64(1 Suppl):S4–16.
[9] Molina S, Castet V, Pichard-Garcia L, et al. Serum-derived hepatitis C virus infection of primary human hepatocytes is tetraspanin CD81 dependent [J]. Journal of Virology. 2008;82:569-574.
[10] Scarselli E, Ansuini H, Cerino R, et al. The human scavenger receptor class B type I is a novel candidate receptor for the hepatitis C virus [J]. The EMBO Journal. 2002;21:5017-5025.
[11] Liu S, Yang W, Shen L, et al. Tight junction proteins claudin-1 and occludin control hepatitis C virus entry and are downregulated during infection to prevent superinfection [J]. Journal of Virology. 2009;83:2011-2014.
[12] Tscherne DM, Jones CT, et al. Time- and temperature-dependent activation of hepatitis C virus for low-pH-triggered entry [J]. J Virol. 2006;80:1734-1741.
[13] Rizzetto M, Canese MG, et al. Immunofluorescence detection of new antigen-antibody system (delta/anti-delta) associated to hepatitis B virus in liver and in serum of HBsAg carriers [J]. Gut. 1977;18(12):997–1003.
[14] Weiner AJ, Choo QL, et al. A single antigenomic open reading frame of the hepatitis delta virus encodes the epitope(s) of both hepatitis delta antigen polypeptides p24 delta and p27 delta [J]. J Virol. 1988;62(2):594–9.
[15] Li J, Wands J. Hepatitis B and D viral receptors [J]. Hepatology. 2016;63(1):11–3.
[16] V.P. Nair, S. Anang, C. Subramani, A. Madhvi, K. Bakshi, A. Srivastava, et al. Endoplasmic Reticulum Stress Induced Synthesis of a Novel Viral Factor Mediates Efficient Replication of Genotype-1 Hepatitis E Virus [J]. PLoS Pathog, 12 (2016), Article e1005521.




