How to Precisely Target Research
Optimal Anti-tag antibody
Anti-tag antibodies and epitope tags can form a stringent antigen-antibody-specific
binding pair. Among these, epitope tags are short known peptides expressed through genetic engineering as
fusions with target proteins. Anti-tag antibodies, serving as highly specific molecular probes, can
recognize these short peptide sequences, thereby enabling qualitative, quantitative, localization, and
affinity purification analyses of the target fusion proteins. In the field of bioengineering, the two
together establish a standardized research platform. As a universal "identifier" fused to target proteins,
epitope tags, in synergy with highly specific anti-tag antibodies, enable high-throughput detection and
purification of various recombinant proteins. This system is widely applied in biopharmaceutical process
monitoring, quality control, and protein interaction analysis in functional genomics, significantly
enhancing research efficiency and standardization.
1. Epitope tag
1.1 Definition
Epitope Tag, also known as tag peptides is a technical tool widely used in molecular biology,
biochemistry, and cell biology research. It refers to the process of linking a known, short exogenous
peptide sequence (typically 8-15 amino acids) to a target protein. N-terminal or C-terminalMan-madeFusion
expression, thereby enabling the detection, purification, or localization of the target protein using
specific antibodies against the tag. Epitope tags and fusion proteins.Significantly distinct:Fusion
protein is a broad concept that refers to a protein expressed by linking two different genes together
through DNA recombination technology (e. g., GFP fluorescent protein + target protein). In contrast, an
epitope tag merely provides a "sequence recognized by an antibody" and does not possess biological
activities such as catalysis or luminescence. Its primary function is to serve as a handle for detection
and purification.
● Common Features of Epitope Tags
- Short in size: Typically composed of 6-15 amino acids. This design
minimally interferes with the spatial conformation of the target protein, making it unlikely to affect
the folding, localization, or biological activity of the target protein.
- Extremely high specificity: These sequences are derived from
exogenous sources such as the influenza virus (HA tag) and human proto-oncogenes (Myc tag), which are
not present in most experimental bacterial or mammalian cells. As a result, they do not produce
cross-reactions, ensuring a clean background.
- Clear immunogenicity: Although short, it can be precisely recognized
by specific monoclonal antibodies with strong affinity.
- Flexible positioning: It can be placed at either end (or even in the
middle) of the protein to accommodate different experimental needs.
1.2 Common Epitope Tags and Their
Characteristics
● Small peptide tag
| Tag Type |
Sequence and Size |
Advantage |
Limitations |
Typical Application Scenarios |
| FLAG tag |
Sequence: DYKDDDDK |
1. High Specificity: Short sequence design reduces non-specific binding, suitable for
experiments such as immunoprecipitation (IP) and Western Blot.
2. AntibodyRich: For example, the M2 anti-FLAG® antibody (Sigma-Aldrich) has been extensively
validated and supports a variety of detection methods.
3. Cleavability: Contains an enterokinase cleavage site, facilitating tag removal to obtain the
native protein.
|
1. Small molecular weight: may affect the native conformation or function of certain small
proteins.
2. Negatively charged: May affect protein migration rate in electrophoresis, requiring
optimization of experimental conditions.
3. Higher cost:AntibodyThe price is higher than some other tag antibodies.
|
1. Protein-Protein Interaction Studies: Co-immunoprecipitation (IP) is used to capture target
proteins and their interacting partners.
2. Drug Screening: Combining flow cytometry (FCM) to analyze changes in cell surface receptor
expression.
3. Gene Therapy: Validation of the Purity and Activity of Viral Vector Packaging Proteins.
|
| Size: 8 amino acids (approximately 1 kDa) |
| HA tag |
Sequence: YPYDVPDYA |
1. Natural Source: Derived from the hemagglutinin protein of the influenza virus, with low
immunogenicity, suitable for live-cell imaging.
2. Multi-application scenarios: Supports Western Blot, IP, flow cytometry (FCM), and
immunofluorescence (IF).
3. High-affinity antibodies: For example, the 12CA5 antibody (Roche) has high sensitivity and
can detect low-abundance proteins.
|
1. Longer sequence: May increase steric hindrance after protein fusion, affecting function.
2. Cross-reactivity Risk: Weak binding with certain proteins in host cells necessitates the
inclusion of negative controls.
3. Low cleavage efficiency: The lack of high-efficiency restriction sites makes tag removal
more difficult.
|
1. Live Cell Dynamic Tracking: Immunofluorescence (IF) for observing protein subcellular
localization (e. g., nucleocytoplasmic transport).
2. Vaccine Development: Detection of surface antigen expression on virus-like particles (VLPs).
3. Cell Signaling Pathways: Analysis of Endocytosis and Degradation of Receptor Tyrosine
Kinases (RTKs).
|
| Size: 9 amino acids (approximately 1 kDa) |
| Myc tag |
Sequence: EQKLISEEDL |
1. Classic Tag: Widely used for protein expression validation, interaction studies, and
subcellular localization analysis.
2. Multi-Technology Compatibility: Supports Western Blot, IP, IF, FCM, and Chromatin
Immunoprecipitation (ChIP).
3. High-sensitivity antibodies: For example, the 9E10 antibody (Santa Cruz) can detect
nanogram-level proteins.
|
1. Larger molecular weight: may affect the folding or activity of small proteins.
2. Charged Sequence: May interfere with the surface charge distribution of the protein,
requiring evaluation of its impact on function.
3. Batch-to-Batch Variation: Some antibodies exhibit inter-batch
differences.DifferenceExperimental conditions need to be optimized.
|
1. Transcription Factor Studies: Chromatin Immunoprecipitation (ChIP) analysis of DNA binding
activity.
2. Protein Stability: Monitor protein degradation rates through pulse-chase experiments.
3. Preclinical testing: ELISA quantitative detection of recombinant protein drug concentration
in serum.
|
| Size: 10 amino acids (approximately 1.2 kDa) |
| V5 tag |
Sequence: GKPIPNPLLGLDST |
1. Long Sequence Advantage: Provides more antigenic epitopes, enhancing antibody binding
specificity.
2. High Expression Stability: Stable expression in mammalian cells, suitable for long-term
studies.
3. Multiple Detection Methods: Supports Western Blot, IP, IF, and ELISA, with low background
signal.
|
1. Large molecular weight: May significantly alter the physical properties of small proteins
(such as mobility, solubility).
2. Few Restriction Sites: Lack of universal restriction sites, requiring customized solutions
for tag cleavage.
3. Limited application scope: Fewer antibody options compared to FLAG/HA tags, requiring
verification of antibody compatibility.
|
1. High-throughput protein expression: Validate recombinant protein expression levels in HEK293
or CHO cells.
2. Viral Vector Research: Assessing the assembly efficiency of adeno-associated virus (AAV)
capsid proteins.
3. Structural Biology: Determining the three-dimensional structure of proteins using X-ray
crystallography or cryo-electron microscopy (requires removal of tags).
|
| Size: 14 amino acids (approximately 1.4 kDa) |
● Affinity purification tag
| Tag Type |
Sequence |
Size |
Advantage |
Limitations |
Typical Application Scenarios |
| 6xHis tag |
HHHHHH (6 histidine residues) |
Approximately 0.8 kDa (excluding the linker peptide) |
1. Small tag, minimal impact on protein structure.
2. Mild purification conditions
3. Low cost and simple operation
|
1. Metal ions may interfere with function.
2. Nonspecific binding
3. Influence on Crystallization or Interaction Studies
|
1. Structural Biology Research (e. g., X-ray Crystallography, Cryo-Electron Microscopy)
2. Protein Interaction Analysis (e. g., Co-IP, Pull-down)
3. High-throughput expression screening (small tags to reduce interference)
|
| GST (Glutathione S-transferase) |
Approximately 211 amino acids (using E.coli GST as an example) |
Approximately 26 kDa |
1. Promote protein solubility and folding
2. The purification steps are simple.
3. The tag is removable.
|
1. The tag is large.
2. Glutathione has a high cost.
3. Fusion proteins are prone to degradation.
|
1. Eukaryotic protein expression (such as mammalian cells or yeast systems)
2. Inclusion Body Protein Refolding (GST Facilitates Correct Folding)
3. Enzyme Activity Study (Functional Analysis After tag Removal)
|
| MBP (Maltose-Binding Protein) |
Approximately 370 amino acids |
Approximately 42.5 kDa |
1. Significantly improve solubility
2. Simple purification steps
3. The tag is removable.
|
1. Large tag
2. The cost of amylose resin is high.
3. The elution conditions may affect the activity.
|
1. Membrane proteins or difficult-to-express proteins (MBP significantly enhances solubility)
2. Functional Studies (e. g., Enzyme Activity, Binding Capacity Assays)
3. Vaccine Development (MBP Fusion Antigen Enhances Immunogenicity)
|
● Fluorescent protein tag
① GFP and its variants
GFP (Green Fluorescent Protein): Derived from jellyfish, composed of 238 amino acids (approximately 27
kDa), emits green fluorescence under blue light excitation (excitation peak 488 nm, emission peak 507 nm).
Variant Optimization:
- EGFP (Enhanced GFP): Through F64L and S65T mutations, fluorescence intensity
is increased by 35-fold, photostability is enhanced, maturation speed is accelerated, and it is suitable
for expression in mammalian cells at 37°C.
- Emerald GFP: Further optimized for enhanced photostability and faster
maturation, ideal for tracking dynamic processes such as cell division.
- Superfolder GFP: Strong acid resistance, high folding efficiency, suitable
for fusion expression prone to inclusion body formation.
- CFP (Cyan Fluorescent Protein): Excitation peak at 433/453 nm, emission peak
at 475/510 nm, commonly used as a FRET donor paired with YFP.
② YFP
YFP (Yellow Fluorescent Protein): The emission spectrum is red-shifted to the yellow region (excitation
peak at 513 nm, emission peak at 527 nm), used to distinguish GFP signals.
Advantage
- Non-invasive real-time imaging: Directly observe protein dynamics in living
cells using fluorescence microscopy without the need for cell fixation or substrate addition.
- High Sensitivity and Low Background: The autofluorescence properties avoid
interference from other intracellular products, making it suitable for high-throughput drug screening.
- Multicolor imaging capability: Combined with variants such as RFP and YFP,
enables simultaneous tracking of multiple targets (e. g., co-localization analysis).
Limitations
- Tagging Effect: May alter the phase separation behavior of the target
protein (e. g., GFP-tagged Dhh1 increases the critical concentration for phase separation),
necessitating validation with unlabeled controls.
- Phototoxicity: Prolonged excitation may cause cellular damage, necessitating
optimization of light dosage (e. g., using LED light sources).
- Spectral Overlap: Some variants (e. g., YFP and GFP) have overlapping
spectra, which limits the resolution of multicolor imaging.
Typical Application Scenarios
- Protein Localization and Dynamic Tracking: tag Tubulin to Observe the Cell
Division Process.
- Gene Expression Report: GFP is placed downstream of the promoter to monitor
gene activation status in real time.
- Protein Interaction ResearchStudy: Detection of Calmodulin Binding to Target
Proteins via FRET Technology (e. g., CFP-YFP Pair).
③ RFP and mCherry
RFP (Red Fluorescent Protein): Cloned from corals (e. g., DsRed with 225 amino acids), emits red
fluorescence (excitation peak at 558 nm, emission peak at 583 nm), but matures slowly and tends to form
tetramers.
mCherry: An optimized variant of DsRed, monomeric form (236 amino acids), excitation peak at 587 nm,
emission peak at 610 nm, with high brightness and photostability. Rapid Maturation, Low background
interference and other advantages, and due toMonomer formYesAvoid interference from protein fusion caused
by tetramers (such as DsRed).
Advantage
- Long-wavelength excitation: Reduces interference from cellular
autofluorescence, suitable for deep tissue imaging.
- Broad adaptability: Already applied in both prokaryotic and eukaryotic cells
(e. g., mammalian cells, yeast).
- Easy to operate: The expression system does not require inducers, making it
suitable for scenarios where inducer addition is difficult (such as recombinant bacteria expressing
proteins in the gastrointestinal tract of experimental animals).
- Compatibility: Can be combined with GFP or other reporter genes to achieve
multicolor labeling and co-localization studies.
Limitations
- Tag Effect: Although it is a monomer, it may still slightly affect the
function of the target protein when expressed in fusion (requires verification with an unlabeled
control).
- Cost: The cost of RFP-related reagents (such as antibodies) is relatively
high.
- Spectral Limitations: Weaker tissue penetration compared to far-red
fluorescent proteins (such as iRFP).
Typical Application Scenarios
- Long-term Tracking: Utilizing the characteristic of strong photostability to
observe dynamic changes in the cell cycle or gene expression.
- Multicolor Imaging: In Combination with GFPDistinguish between different
proteins or cellular structures (e. g., co-localization of endoplasmic reticulum and mitochondria).
- Flow Cytometry and Immunofluorescence: Analyze cell phenotypes or protein
expression levels by detecting or amplifying fluorescent signals using anti-RFP antibodies.
④ Comprehensive Applications of Fluorescent Protein Tags in Live Cell
Imaging
Multicolor Imaging Strategy
- Combination Selection: GFP (blue channel) and mCherry (red channel) are
paired to avoid spectral overlap, enabling high-resolution co-localization analysis.
- Application Case: Simultaneously labeling microtubule protein (GFP) and
mitochondria (mCherry) to observe changes in mitochondrial distribution during cell division.
Dynamic Process Monitoring
- FRAP (Fluorescence Recovery After Photobleaching): tag GFP fusion proteins
and analyze protein transport dynamics by measuring the fluorescence recovery rate in the bleached
region.
- FRET Technology: Utilizing CFP-YFP Combinations to Detect Protein
Interactions (e. g., Changes in Calcium Ion Concentration).
Deep Tissue Imaging
- Two-photon microscopy: Utilizes long-wavelength excitation for red
fluorescent proteins (e. g., mCherry), reducing light scattering and enabling deep tissue imaging (e.
g., brain slices).
- Near-infrared fluorescent protein (iRFP): Emission wavelength 650-900 nm,
with stronger tissue penetration, suitable for in vivo imaging (e. g., tumor models).
Super-resolution imaging technology
- STED Microscopy: Utilizing differences in fluorescence lifetimes of
fluorescent proteins to surpass the diffraction limit and resolve internal substructures of condensates
(such as Huntingtin protein aggregates).
- Single-Molecule Localization Microscopy (SMLM): Achieves nanoscale
resolution imaging through photoactivation or photoconversion of proteins (such as PA-GFP, Kaede).
It is worth mentioning that beyond the direct use of fluorescent proteins, protein
tagging technologies such as SNAP-tag and HaloTag are also continuously evolving. These technologies can
specifically bind to exogenous fluorescent dyes, combining the targeting capabilities of genetic encoding
with the superior optical properties of chemical dyes, thereby offering more options for live-cell
imaging.
CUSABIO offers a comprehensive
range of tag antibodies, including 6*His, GFP, c-MYC, GST, E-Tag, HA-Tag, Flag Tag, and Sumo Tag. These
antibodies are purified via antigen affinity purification, enabling specific binding to correspondingly
tagged proteins for the identification and purification of target proteins. With a complete variety and
high specificity, they ensure the smooth progress of your experiments.
2. Anti-tag antibodyEpitope tagRecognition mechanism
Anti-tag antibodies are highly specific antibodies developed specifically for particular epitope tags.
Their mechanism of action is based on the classic principle of antigen-antibody specific recognition..In
the recognition mechanism of tag-antibody binding, anti-tag antibodies primarily function through
three-dimensional conformational recognition rather than simply reading the primary sequence. Although
epitope tags are linear amino acid sequences, they fold into specific secondary structures (such as
helices or turns) on the surface of the target protein. The antigen-binding fragment (Fab) of the antibody
recognizes their three-dimensional spatial shape and surface charge distribution. This recognition does
not depend on the function of the target protein itself but solely on whether the tag is exposed.
Additionally, to avoid false positives, anti-tag antibodies typically exhibit extremely high sequence
specificity. The synergy of these two aspects enables researchers to use the same set of tools to study
any tagged protein without the need to develop specific antibodies for each target protein individually.
Detection (e. g., Western Blot), visualization and localization (e. g., immunofluorescence), and
affinity purification (e. g., immunoprecipitation). significantly enhancing the standardization and
efficiency of the experiments.
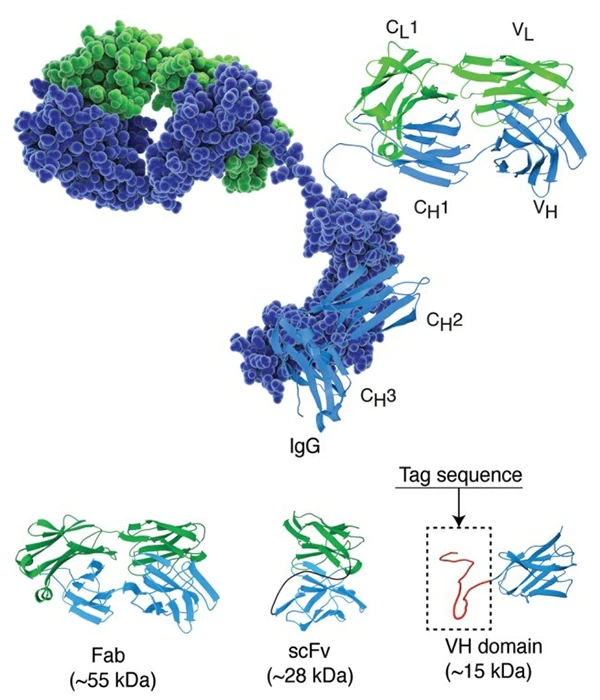
Image source: PMID: 25426869
The structure of immunoglobulin G in space-filling and
cartoon representation with light and heavy chain colored in green and blue, respectively. Lower panel
illustrates crystal structures of the antibody fragments FAB (fragment antigen-binding), scFv
(single-chain fragment variable) and single domain antibody. A peptide tag of approximately 30 residues
is illustrated at the c-terminal of the domain antibody. The immunoglobulin structure used in this
figure is based on the RCSB Protein Data Bank entry 1igt.
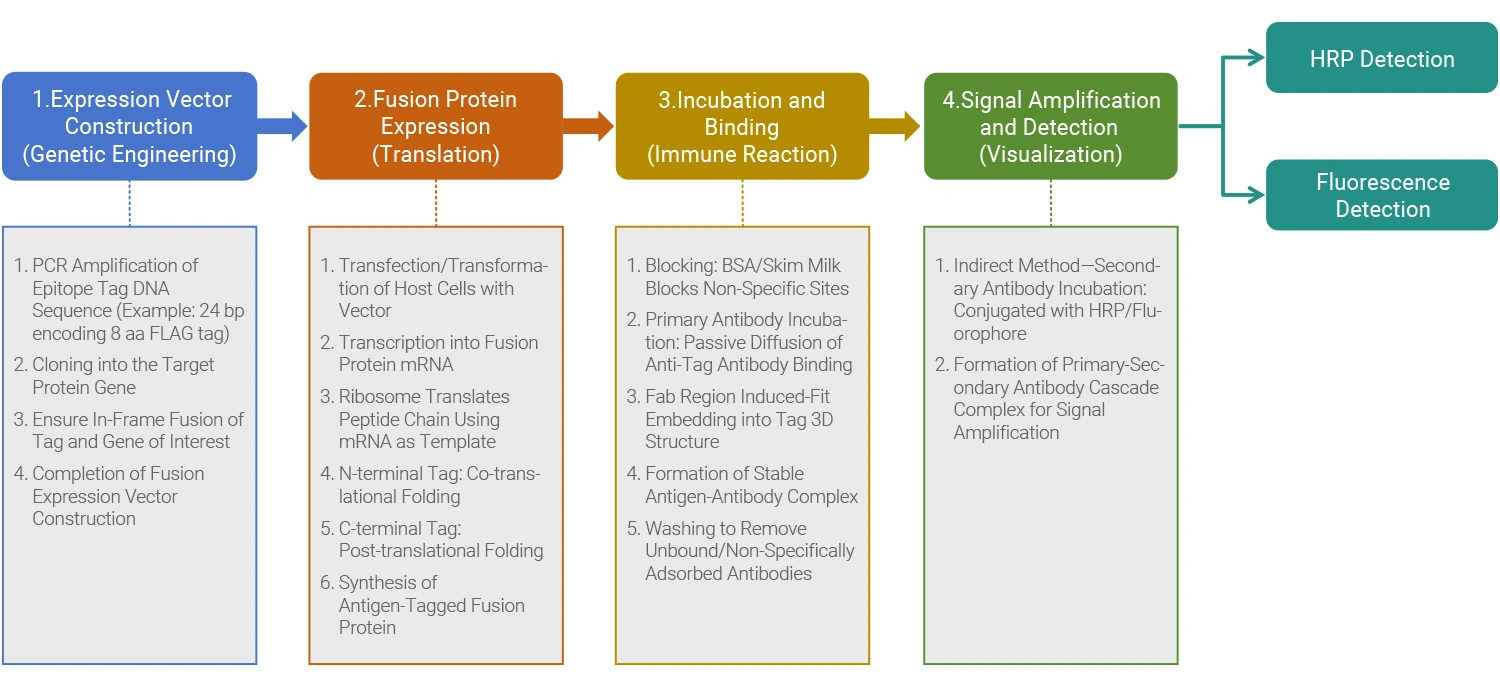
● SpecificTesting Process
From Marker Protein Expression to Antibody Recognition, Usually divided intoFour key steps:
3. Anti-tag antibody vs. Protein-specific antibody
| Comparison Dimensions |
Anti-tag antibody |
Protein-specific antibody |
| Development Goals |
For standardized tag sequences (such as FLAG, His) development |
Unique epitopes specific to a particular natural protein |
| Development Cycle and Cost |
Short development cycle (2-3 weeks), low cost, relies on hybridoma technology or recombinant
expression platforms, with high batch-to-batch consistency. |
Long development cycle (several months), high cost, and requires multiple rounds of validation
for specificity. |
| Scope of Application |
Any fusion protein (universal tool) bearing this tag |
Recognize only specific natural proteins (specialized tool) |
| Detection Principle |
- Exogenous Protein Detection: Validating the expression of
recombinant proteins after transfection or infection.
- Indirect detection: Requires genetic engineering to fuse a tag to
the target protein for use, detecting the tag rather than the protein itself.
|
- Endogenous Protein Detection: Investigating the expression levels
and localization of native proteins in tissues or cells.
- Direct Detection: Directly identifies the unique epitopes of
natural proteins without the need for genetic modification of the sample.
|
| Application Scenarios |
Universal TestingPurification: Suitable for Western Blot, immunofluorescence,
immunoprecipitation, affinity chromatography, etc. Supports high-throughput experiments and
industrial production (e. g., antibody drug purification, enzyme preparation production,
recombinant protein expression monitoring, protein interaction preliminary screening). |
Specificity Studies: Suitable for scenarios such as protein-protein interaction network
analysis, signaling pathway research, disease-related protein detection, and native conformation
analysis, requiring differentiation between endogenous and exogenous proteins. |
| Core Advantages |
- High versatility: A single antibody can be used to detect and
capture hundreds of different tagged proteins.
- High signal-to-noise ratio: Typically no endogenous background in
mammalian cells (e. g., FLAG/Myc tags).
- Reagent Stability: Minimal batch-to-batch variation ensures highly
standardized experimental results.
- Gentle Elution: Specific tags (such as FLAG) can be eluted using
peptide competition, preserving protein activity.
|
- Physiological Relevance: Accurately reflects the expression level
and localization of proteins in their native state.
- No genetic modification required: directly applicable to wild-type
animals, clinical, or pathological samples.
- Recognition of Modification States: Capable of distinguishing
between different splice variants or post-translational modifications of proteins.
- Versatile Applications: Suitable for various detection platforms
such as immunohistochemistry, flow cytometry, and ELISA.
|
| Limitations |
- Tag Dependency: May be influenced by tag exposure level, position,
or modifications (e. g.,N-terminal/C-terminal differences).
- Functional Interference: Tags may affect the native conformation
or function of the protein.
- Environmental Sensitivity: May fail under extreme conditions (e.
g., high imidazole concentration, strong acids or bases).
|
- Long development cycle: Requires several months and high costs.
- Narrow scope of application: only targets specific proteins.
- Cross-reactivity risk: There may be non-specific binding.
- High demand for experimental optimization: Conditions need to be
adjusted to reduce background interference (e. g., sealing and washing steps).
|
4. Why are anti-tag antibodies used in research?
Anti-tag antibodyAsA specific antibody targeting artificially introduced short peptide sequences, known
as "tags," has become an extremely versatile and powerful tool in biomedical research. Its core advantage
lies in the fact that researchers no longer need to develop and validate specific antibodies from scratch
for each target protein. Instead, they can utilize existing, highly optimized anti-tag antibody systems to
detect, purify, localize, and functionally manipulate a wide variety of proteins. The following sections
will elaborate on its applications and core advantages.
4.1 Anti-tag antibodies are primarily
used in scenarios such as:
● Protein Crystallization and Structure Determination
In structural biology, obtaining high-quality protein crystals is a significant challenge. Anti-tag
antibodies can serve as "crystallization chaperones" to facilitate the crystallization of target proteins.
For example, the Fab fragment of the monoclonal antibody NZ-1, which recognizes the PA tag, can form a
complex with the target protein containing the inserted PA tag, thereby promoting co-crystallization
[1]. This method circumvents the difficulty of developing conformation-specific antibodies for
each target protein by inserting a tag into specific loop regions of the target protein (such as β-hairpin
structures). It leverages the high-affinity binding of antibodies to the tag to stabilize the protein
conformation, ultimately yielding crystals suitable for high-resolution structural analysis.
● Protein Detection and Visualization
- Western Bllot (Western Blot) and Immunofluorescence: For Detection
and Quantification of Exogenously Expressed Tagged Proteins.
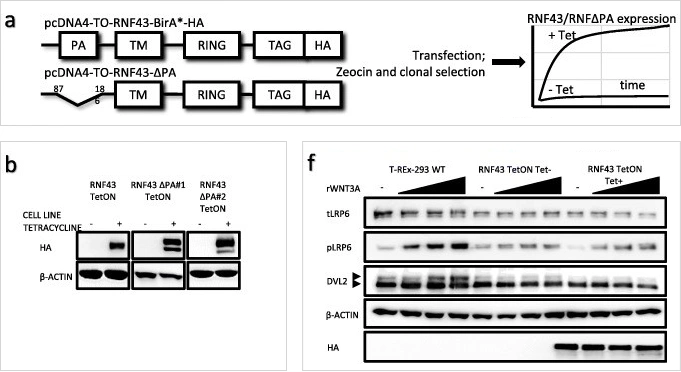
Image source: PMID: 30666745
Corresponding English translation:Inducible RNF43 and
RNF43ΔPA expression in T-REx 293 cell line.
a. Schematic representation of the used experimental models.
b. Western blot showing Tet-induced expression of HA-tagged RNF43 constructs in stable sell lines.
f. Western blot analysis of canonical Wnt pathway activation by application of the increasing
concentrations of rWNT3A (40, 60 and 100 ng/mL) after 3 h of treatment.
- Immunoelectron Microscopy: Used for localizing membrane protein
complexes at the ultrastructural level. This is achieved through engineered antibody fragments (Fv),
where one end binds to the epitope of the target protein, and the other end carries a tag (such as a
Strep-tag or c-myc tag). High-resolution localization is then accomplished using corresponding detection
systems, such as colloidal gold-labeled streptavidin or anti-tag antibodies.
- Immunohistochemistry/Cytochemistry: The basic principles are the same
as above, used to localize tagged proteins in tissue or cell sections.
● Chromatin Immunoprecipitation
Chromatin immunoprecipitation is a key technique for studying protein-DNA interactions in vivo. When
specific antibodies against endogenous transcription factors are unavailable or ineffective, expressing
exogenous epitope-tagged fusion proteins and performing immunoprecipitation with highly specific anti-tag
antibodies serves as a reliable and efficient alternative [2]. This method avoids the
time-consuming and labor-intensive preparation of transgenic stable lines; for example, transient
expression of tagged proteins in Arabidopsis protoplasts allows ChIP experiments to be completed within
four days.
● Protein purification
Anti-tag antibodies can be used for rapid, one-step affinity purification of recombinant proteins. For
example, Sanae Tabata.Waiting for someone in 2010.Using the anti-TARGET tag antibody P20.1, a sensitive
screening system can be established to rapidly select high-yield cell lines from mammalian cell culture
supernatants and standardize the purification of milligram-scale target proteins. Similarly, in the
production of Fv fragments, one-step affinity purification can be achieved through their fused Strep tag.
● Functional Research and Regulation
Anti-tag antibodies can not only "see" and "grab" but also "interfere with" and "activate" protein
functions.
- Functional Interference: The binding of certain anti-tag antibodies
can block protein-protein interactions. For example, an antibody against the SV40 large T antigen, whose
epitope overlaps with the site where T antigen binds to RPA70N, can directly inhibit the interaction
between the two.
- Function Activation: In synthetic biology applications, anti-tag
antibodies can be used for artificially induced protein dimerization to activate signaling. For example,
a transmembrane fusion protein expressed in protoplasts, with a tag at its extracellular end, can
undergo conditional dimerization and activation of an intracellular reporter enzyme (GUS) upon external
addition of the corresponding anti-tag antibody. This enables signal amplification and detection.
● Diagnosis and Detection Technology Development
The anti-tag antibody system provides modular tools for constructing highly sensitive detection
platforms.
- Digital ELISA and Biosensors: The prototype vesicle-based array
utilizes transmembrane signaling induced by anti-tag antibodies to achieve highly sensitive, digital
detection of specific antibodies (such as trastuzumab) and even small molecule antigens (such as
caffeine), while reducing cumbersome washing steps.
- Magnetic Relaxation Switch: In magnetic particle-based sensing, the
detection sensitivity can be significantly enhanced by increasing the target valency through the use of
anti-tag antibodies and their secondary antibodies.
- Protein Chip: In self-assembling protein chip technology, anti-tag
antibodies are first printed onto the chip. Subsequently, a cell-free expression system is used to
synthesize tag-fusion proteins in situ on the chip, which are immediately captured by the antibodies,
enabling high-throughput production of functional protein microarrays.
● Clinical Diagnosis and Treatment Applications
In the clinical field, radionuclide-labeled anti-tag antibodies can be used for tumor localization and
surgical navigation. For example, in recurrent colorectal cancer surgery, the use of iodine-125-labeled
anti-tag antibodies (CC49 monoclonal antibodies) for radioimmunoguided surgery can detect more occult
lesions that traditional imaging and surgical exploration cannot identify, thereby altering surgical plans
and improving tumor resection rates.
4.2 Core Advantages
Based on the above scenarios, the advantages of anti-tag antibodies are primarily reflected in the
following aspects:
- Versatility and Efficiency: A well-established system of anti-tag
antibodies (such as anti-HA, myc, FLAG, and PA tags) can be applied to any protein labeled with the
corresponding tag, eliminating the need to customize antibodies for each new protein, thereby
significantly saving time and costs.
- High Specificity and High Affinity: Many commercially available
anti-tag antibodies are meticulously screened and optimized, offering exceptionally high specificity and
affinity.
- Flexibility: Tags can be relatively freely inserted into the
N-terminus, C-terminus, or internal loop regions of the target protein to accommodate various
experimental needs, such as surface display or avoiding interference with functional domains.
- Improving Experimental Success Rate and Reproducibility: Using
validated anti-tag antibodies for ChIP, purification, or detection can prevent experimental failures
caused by poor-quality endogenous antibodies, resulting in more stable and reliable outcomes.
- Achieving Innovative Experimental Design: Anti-tag antibodies provide
critical components for cutting-edge technologies, such as constructing protein chips in cell-free
expression systems, building signal transduction systems in synthetic vesicles, and developing novel
biosensors, thereby advancing methodological progress.
5. Emerging Tag Technology
5.1 Next-Generation Small Tags HiBiT
and Other Ultra-Small Tags
Traditional protein tags (such as GFP and HaloTag), while powerful, have relatively large molecular
weights (typically over 20 kDa), which may interfere with the native structure, function, localization,
and interactions of the protein of interest (POI). The new generation of ultra-small tag technologies,
represented by the HiBiT peptide, is addressing these challenges.
The core advantages of the HiBiT tag lie in its extremely small size and its highly luminescent
complementary system. HiBiT is a short peptide composed of only 11 amino acids (approximately 1.3 kDa). It
does not emit light on its own but can bind with high affinity to a larger complementary protein fragment
(LgBiT, 18 kDa) to form a complete and active NanoLuc luciferase. This design offers multiple
revolutionary advantages:
- Minimize interference with the target protein: Due to its extremely
small size, fusing the HiBiT tag to the N- or C-terminus of the target protein causes significantly less
interference with the protein's native folding, membrane localization, signal transduction, and
protein-protein interactions compared to traditional large tags. This enables the study of proteins in a
nearly native physiological environment, making the data more physiologically relevant [3].
- Achieving High Signal-to-Noise Ratio for Selective Detection of Cell
Surfaces: The LgBiT protein cannot penetrate the cell membrane. Therefore, luminescence signals
are generated only when the HiBiT tag is located on the surface of living cells and complements the
externally added LgBiT. This characteristic is ingeniously utilized for real-time, dynamic monitoring of
changes in the quantity of cell surface receptors, such as the internalization process of G
protein-coupled receptors (GPCRs) following ligand stimulation. This method exhibits an extremely low
background because intracellular receptors cannot produce signals [4].
- Ultra-high sensitivity and wide dynamic range: NanoLuc luciferase is
one of the brightest known luciferases. The HiBiT/LgBiT complementation system inherits this high
brightness characteristic, enabling the detection of extremely low-abundance target proteins, even in
research models with minimal exogenous protein expression. Its wide dynamic range facilitates precise
quantification of signals ranging from weak to strong [4].
- Supports multiple detection and dynamic process analysis: The HiBiT
system is compatible with other Tag technologies (such as HaloTag and SNAP-tag), enabling multicolor
single-molecule imaging and simultaneous tracking of multiple protein dynamics. Additionally, the
HiBiT-based secretion reporter system allows for real-time monitoring of dynamic biological processes,
such as the bacterial type III secretion system, with high sensitivity and throughput. [5].
- Simplifying Virology and Drug Screening Research: The HiBiT tag can
be inserted into specific sites of viral proteins, such as the ORF2 protein of hepatitis E virus, to
construct infectious viruses carrying a luminescent reporter gene. This recombinant virus enables
convenient and quantitative real-time monitoring of viral infection and replication processes, greatly
facilitating the evaluation of antiviral drugs and neutralizing antibodies [6].
5.2 Tags Recognizable by Nanobodies
The working principle of the nanobody-tag system involves camelid-derived or artificially designed
single-domain antibodies that can bind with high affinity and specificity to short peptide sequences of
6–14 amino acids. These short peptide tags can be fused to the N-terminus or C-terminus of target
proteins. After expression, the nanobodies serve as versatile tools for protein capture, visualization, or
manipulation.
● Nanobody Recognizable Tag Types
- High-specificity short peptide tag: ① 6-7 amino acid short peptide
tag: A high-efficiency purification system based on nanobodies enables one-step purification of various
soluble and membrane proteins with high purity and recovery rates. Bound proteins can be gently eluted
using competitive synthetic peptides, achieving recovery rates exceeding 90%. ② 14-mer peptide epitope
tag (e. g., Nb6E system): Nanobody Nb6E interacts with a 14-amino acid peptide epitope, identifying key
residues and proposing an interaction model to enhance the potency of engineered adenosine A2A receptor
ligands [7].
- Tag system for covalent cross-linking: Synthetic peptide epitopes
paired with nanobodies enable rapid and specific covalent cross-linking on the surface of living cells.
The cross-linking reaction proceeds at a fast rate, making it suitable for tag live cells and studying
the effects of ligand cross-linking on receptor signaling.
- Tag Systems for Protein Localization and Proximity tag: ALFA tag
System, Based on ALFA Tag - Nanobody Pairing for Protein Localization and Proximity Proteomics in
Mycobacteria, Applicable to Various Microbial Systems.
- C-terminal recognition sequence for secretion production: The LipC
secretion system of Serratia marcescens recognizes the C-terminal Val-Thr-Val sequence of nanobodies,
enabling the one-step secretion of nanobodies to the extracellular environment, thereby constituting a
secretion recognition tag based on a specific sequence.
● Advantages of Nanobody-Tag Systems
- High Affinity and Specificity: Binding at sub-nanomolar to picomolar
levels with minimal background interference.
- Minimal tag: Minimal impact on the structure and function of the
target protein.
- Gentle Elution: Helps Maintain Target Protein Activity
5.3 TEV Protease Cleavage Site
Although the interference from small tags and nanobodies has been minimized, in certain detailed studies,
it is still necessary to completely remove the tags to obtain protein products that are entirely
consistent with their native state. Cleavable tag systems have emerged to address this need, with the TEV
protease cleavage site being one of the most commonly used and efficient systems.
TEV protease is a highly specific protease that recognizes its unique
seven-amino-acid sequence (Glu-Asn-Leu-Tyr-Phe-Gln-Gly or similar variants) and cleaves between Gln and
Gly/Ser. In protein purification or functional research workflows, the decision to introduce and
ultimately remove tags is primarily based on the following considerations:
- Obtain un-tag purified protein: In recombinant protein expression,
larger affinity tags (such as His-tag, GST-tag) are commonly used to simplify the purification process.
By introducing a TEV protease cleavage site between the affinity tag and the target protein, high-purity
tag-free target protein can be obtained through enzymatic cleavage after purification.
- Control the activation or release of proteins: In synthetic biology
or cellular engineering, cleavable tags can be used to design conditionally activated proteins. For
example, a functional domain or inhibitory peptide can be linked to the main protein via a TEV site,
rendering it inactive. Only under specific conditions (such as the expression or addition of TEV
protease) does cleavage occur, releasing the functional domain and activating the protein.
- Addressing Special Requirements in Research Models: In Tag Antibody
Design a cleavable linker between the target viral protein and the reporter gene, allowing the removal
of the reporter gene after preliminary studies involving reporter gene detection, thereby obtaining the
viral protein without the reporter gene for more in-depth functional validation.
6. How to Choose the Appropriate Anti-Tag Antibody
In bioengineering experiments, anti-tag antibodies are crucial tools that link target proteins with
detection technologies. However, with the wide array of tag antibody products available on the market, how
can one accurately select the right one based on experimental needs? We can From the experimental
objective From tag and Tag Design to Antibody Characteristics and Technical Compatibility, Systematically
Organize the Selection Strategy for Anti-Tag Antibodies to Help You Improve Experimental Efficiency and
Data Reliability.
6.1 Selecting tag antibodies based on
experimental objectives
● Protein Localization Studies: Fluorescent or Small Peptide
Tags Are Preferred
For protein localization studies, it is essential to visually present the subcellular distribution of
target proteins through fluorescent signals or immunostaining. In such cases, fluorescent protein tags
(such as GFP, mCherry) or small peptide tags (such as HA, FLAG) are more suitable.
Fluorescent tag: Directly provides visual signals without additional staining
steps, suitable for live-cell imaging. The microscope excitation light source needs to be matched (e. g.,
GFP requires a 488 nm laser).
Small Peptide Tag: Compatible with fixed cell samples via immunofluorescence
or immunohistochemistry detection, featuring a small tag size and minimal interference with protein
function.
● Protein purification
Protein purification is achieved by efficiently capturing the target protein through the high-affinity
binding of a tag to a ligand, followed by its release under specific elution conditions, such as pH
adjustment or the use of competitive agents. NeedSelect tags based on the characteristics of the target
protein (e. g., prioritize His tags for membrane proteins, prioritize MBP tags for proteins prone to
aggregation). AndAfter purification, the tag must be removed by enzymatic digestion or chemical cleavage
to avoid interference with downstream functional experiments.
Common tags include His, GST, and MBP tags.
His-tag: Suitable for immobilized metal affinity chromatography (IMAC), with
mild elution conditions (e. g., imidazole gradient), but attention should be paid to interference from
endogenous histidine in host proteins.
GST tag: Eluted via glutathione, suitable for large-scale purification, but
the tag is relatively large (26 kDa) may affect protein solubility.
MBP tag: Enhances solubility of target proteins, eluted with maltose, suitable
for difficult-to-fold proteins.
● Protein-Protein Interaction Studies
Interaction studies require the capture of protein complexes through co-immunoprecipitation (Co-IP) or
pull-down techniques, and the tags must meet the following criteria:
Small size: Avoids steric hindrance that could interfere with natural
interactions (e. g., FLAG, HA, and Myc tags consist of only 8–10 amino acids).
High specificity: Antibodies must strictly recognize Do not tag sequences to
avoid cross-reactions.
FLAG, HA, Myc tagsAvailableMinimize interference.
6.2 Tag Size and Position
● The Impact of Tag Size on Protein Function
The molecular weight of the tag is recommended not to exceed 10% of the target protein (e. g., 50 kDa
protein preferentially selects <5 kDa tag). A tag that is too large may interfere with the folding,
stability, or interaction interface of the target protein. During the practical operation process,Predict
the interface between the tag and the target protein using AlphaFold to avoid covering critical functional
domains.
● N-terminal vs. C-terminal tag
N-terminal tag: Suitable for cytoplasmic proteins without signal peptides, but
may interfere with the transport of secreted proteins (e. g., N-terminal tag of antibody light chains can
block secretion).
C-terminal tag: More versatile, but care must be taken to avoid disrupting
localization signals at the C-terminus (such as mitochondrial targeting sequences).
Internal tag: Tags can be inserted into flexible loop regions of long
proteins, requiring screening of mutation libraries to identify sites that do not affect function.
Internal tag is suitable for functional key regions at the N/C-terminus of the target protein (such as the
enzyme active center).
6.3 Antibody Characteristics:
Specificity, Sensitivity, and Validation Data
● Monoclonal vs. Polyclonal Antibodies
Monoclonal antibodies: High specificity, minimal batch-to-batch variation,
suitable for quantitative experiments (e. g., WB, ELISA), but with high development costs.
Polyclonal antibodies: High sensitivity, capable of recognizing multiple
epitopes, suitable for detecting low-abundance proteins, but require strict quality control for
batch-to-batch variability.
For critical experiments (such as clinical sample testing), monoclonal antibodies are prioritized.
Polyclonal antibodies can be used during preliminary experiments or screening stages to reduce costs.
● Sensitivity and Specificity
Sensitivity: Select antibodies based on the expression level of the target
protein (e. g., high-affinity antibodies are required for low-expression proteins, such as those with KD
< 1 nM).
Specificity: Detect cross-reactivity of the target protein with host proteins
(e. g., E.coli, HEK293) via Western blot (WB), prioritizing antibodies validated using knockout cell
lines.
● Application Verification
Validation Data: Verify that the antibody has been validated across multiple
techniques such as WB, IP, IF, and ChIP, and avoid using antibodies validated solely by ELISA for IP
experiments.
● Host Species Selection
Primary antibody host: Commonly used hosts include rabbit, mouse, and goat.
The primary antibody host species must match the secondary antibody host species (e. g., rabbit anti-FLAG
primary antibody should be paired with anti-rabbit IgG secondary antibody). To avoid cross-binding of
secondary antibodies, select primary antibodies from different host sources (e. g., mouse anti-HA + rabbit
anti-Myc).
6.4 Technical Compatibility
● Fixation Method Compatibility: Formaldehyde vs. Methanol
Formaldehyde Fixation: Cross-links proteins, potentially masking linear
epitopes, necessitating the selection of antibodies that recognize conformational epitopes (such as
certain fluorescently tag antibodies).
Methanol fixation: Suitable for membrane proteins, but may disrupt the
antigenicity of certain tags (e. g., GST tags are prone to denaturation in methanol).
● Antigen Retrieval Requirements:
Heat-Induced vs. Enzymatic Methods
Heat-induced epitope retrieval (HIER): Suitable for formalin-fixed,
paraffin-embedded sections, where epitopes are restored through high-temperature heating (e. g., 100°C for
20 minutes).
Enzyme Repair: Use Proteinase K or pepsin, optimizing concentration and time
to avoid over-digestion.
● Natural vs. Denatured Protein
Identification (Conformational epitope vs. linear epitope)
Natural Proteins: Antibodies that recognize conformational epitopes should be
selected (e. g., certain fluorescently tag antibodies only bind when the protein is correctly folded).
Denatured protein: Suitable for WB experiments, it is necessary to select
antibodies that recognize linear epitopes (such as His-tag antibodies, which can still bind in SDS-PAGE).
6.5 Series Tag Strategy
● Synergistic Effects of Multiple Identical Tags
For exampleInsert a FLAG tag at each end of the target protein (FLAG-protein-FLAG) to enhance detection
signals via a dual-antibody sandwich assay. Tandem His6 tags (His6-His6) increase binding capacity to
nickel columns, making them suitable for purifying low-expression proteins.
● Combined Application of Different Tags
Purification + Detection Tags: For example, the His tag is used for
purification, and the V5 tag is used for WB detection, to avoid interference from purification tags in
downstream analysis.
Multicolor experimental tags: such as GFP (localization) + HA (interaction) +
Myc (expression level), enabling multidimensional information acquisition.
CUSABIO offers high-quality products covering all scenarios. Rigorously
validated across multiple scenarios.
Now available Free download complete Tag Antibody Product Rapid Selection, Facilitating Smooth
Progress in Scientific Research!
7. Best Practices in Experimental Design
7.1 Design of Tag Fusion Proteins
● Principles for Tag Insertion Positions
Objective: Balance protein function, antibody accessibility, and
structural integrity.
Operating Steps:
- Functional Domain Analysis: Use UniProt, PDB databases, or AlphaFold to
predict the domains and signal sequences of the target protein.
- Antibody Binding Accessibility: Prioritize inserting tags into flexible
regions on the protein surface, such as disordered or random coil regions. Use PyMOL or ChimeraX to
visualize the protein structure and mark potential insertion sites.
- Signal Sequence Evasion: Secretory proteins (such as antibodies) require the
retention of the N-terminal signal peptide, and tags should be inserted downstream of the signal peptide
(e. g., a His tag following the N-terminal signal peptide of the IgG light chain).
● Using Adapter Sequences
Objective: Minimize interference of tags on the conformation of the
target protein and enhance antibody binding efficiency.
Operating Steps:
- Flexible Joint Addition Conditions: Direct linkage of the tag to the target
protein may cause steric hindrance (e. g., when the FLAG tag is adjacent to the transmembrane domain of
a membrane protein).
- Joint Length and Composition Optimization:
Length: Typically, 4-20 amino acids, short linkers (such as GGGGS) are
suitable for small tags (like His), while long linkers (such as (GGGGS)₃) are suitable for large tags
(like GST).
Composition: Avoid proline-rich (Pro) or charged amino acids (such as
Arg, Lys), and recommend using flexible combinations of glycine (Gly) and serine (Ser).
● Expression System Considerations
Objective: Select an expression system based on protein complexity,
balancing yield with functional correctness.
Operating Steps:
- Bacteria vs. Mammalian Expression:
Bacterial expression (e. g., E.coli): Suitable for simple proteins
without post-translational modifications (e. g., His-ERK2), low cost but prone to forming inclusion
bodies.
Mammalian expression (e. g., HEK293): Suitable for proteins requiring
glycosylation or disulfide bonds (e. g., membrane receptor-GFP fusions), but with higher costs.
- Tag Removal Options:
Enzyme Cleavage Site Design: Insert TEV or Thrombin recognition sequences
(e. g., ENLYFQG or LVPRGS) between the tag and the target protein, allowing for tag removal via
enzymatic cleavage after purification.
Self-cleaving tags: Utilize intein tags (such as the IMPACT system) to
achieve auto-cleavage induced by pH or temperature.
7.2 Set appropriate controls
● Negative Control
Objective: Eliminate non-specific background signals.
Operating Steps:
- Untransfected cells: To detect the background fluorescence of the cells or
non-specific binding of antibodies (e. g., autofluorescence of untransfected cells in flow cytometry).
- Empty vector control: Transfect with a vector lacking the target protein (e.
g., pcDNA3.1 empty vector) to verify no cross-reactivity of the tag antibody.
● Isotype Control
Objective: Correct for antibody nonspecific binding (especially in
IF and flow cytometry).
Operating Steps:
- Isotype Control Selection: An antibody from the same species and isotype but
targeting an irrelevant antigen (e. g., for an anti-FLAG primary antibody of rabbit IgG, use a rabbit
IgG isotype-matched control antibody).
- Concentration matching: The concentration of the isotype control should be
the same as that of the primary antibody (e. g., 1 μg/mL).
● Positive Control
Objective: Verify the effectiveness of the experimental system.
Operating Steps:
- Known Marker Proteins: Use validated marker proteins (such as commercially
available GFP-actin fusion proteins) as positive controls for WB or IF.
- Commercial Control Lysate: Use cell lysates containing the target protein
(e. g., HeLa cell lysate for detecting endogenous ERK2).
8. Common Issues Troubleshooting
| Fault Type |
Possible reasons |
Phenomenon Analysis |
Solution |
| No signal or weak signal |
Low expression level of the marker protein. |
Low expression level of the marker protein. |
Verification of Transfection/Transduction Efficiency: Use fluorescent co-transfection or flow
cytometry to confirm the proportion of positive cells. |
| Optimization conditions: Conduct induction time/concentration gradient experiments and replace
the promoter. |
| Check protein stability: Treat cells with proteasome inhibitors (such as MG132) and observe
whether the signal is restored. |
| Insufficient Antibody Concentration |
The positive control shows a weak signal, while the sample shows no signal. |
Titration experiment: Incubate the same sample membrane with different dilutions (e. g., 1: 500,
1: 1000, 1: 5000) to determine the optimal concentration. |
| Recommended starting dilution: For new antibodies, begin with the middle value of the range
specified in the instructions (e. g., if the range is 1: 100-1000, use 1: 500). |
| Tag Epitope Masked |
The tag antibody shows no signal, but the target protein-specific antibody shows a
signal. |
Protein Folding: Increase the sample boiling temperature (95°C, 10 min) or enhance the strength
of the denaturant to fully expose the epitope. |
| Alternative tag placement: If the N-terminal tag is blocked, attempt to construct a plasmid with
a C-terminal tag. |
| Transfer Efficiency Issues |
There is residual protein in the gel staining, or the markers on the membrane are
unclear. |
Protein size consideration: Large molecular proteins (>150 kDa) Choose wet transfer, low
methanol, long time; small molecular weight proteins (<20 kDa) Use a 0.2 μm pore size membrane
for a short duration.
|
| Transfer Optimization: After transfer, routinely perform Ponceau S staining to verify the
transfer efficiency and observe loading consistency. |
| High Background Signal |
Antibody Concentration Too High |
The entire membrane appears blurred, with no clear bands visible. |
Optimization Strategy: Reduce the concentration of the primary or secondary antibody, and
shorten the incubation time. Pay special attention to the concentration of the secondary antibody,
as it is often the main source of high background. |
| Inadequate Sealing |
The background is uniform and has a matte texture. |
Blocking Buffer Selection: Typically use 5% skim milk; switch to 5% BSA when detecting
phosphorylated proteins (to avoid interference from casein in milk). |
| Blocking Buffer Selection: Typically use 5% skim milk; switch to 5% BSA when detecting
phosphorylated proteins (to avoid interference from casein in milk). |
| Inadequate Washing |
The background shows localized patchy or streaky stains. |
Wash Buffer Composition: TBST (containing Tween 20) is routinely used. |
| Washing Duration and Frequency: Emphasize "small amounts, multiple times," with each session
lasting 5-10 minutes, repeated more than 5 times. |
| Non-specific Binding |
In addition to the target band, there is also a noticeable background of unintended
bands. |
Antibody Quality Issues: Use affinity-purified antibodies or monoclonal antibodies validated in
the literature. |
| Buffer Optimization: Increase the salt concentration or Tween-20 content in the antibody
dilution/wash buffer to reduce hydrophobic interactions. |
| Multiple bands detected |
Post-translational Modification |
Bands larger than the target protein appear, or multiple bands with a regular
pattern. |
Expected modifications: Review literature to confirm known modifications. Perform
de-modification treatments (e. g., dephosphorylation enzymes) as control verification. |
| Mobility Shift: Modifications such as phosphorylation/glycosylation can alter protein migration
rates, which is a normal phenomenon. |
| Post-translational Modification |
Bands larger than the target protein appear, or multiple bands with a regular
pattern. |
Expected modifications: Review literature to confirm known modifications. Perform
de-modification treatments (e. g., dephosphorylation enzymes) as control verification. |
| Mobility Shift: Modifications such as phosphorylation/glycosylation can alter protein migration
rates, which is a normal phenomenon. |
| Variability in Myc Tag Recognition |
There are many bands, even the internal reference channel has background bands. |
Antibody Concentration Adjustment: Reduce the antibody concentration and perform a titration
experiment. |
| Additional Control: Set up knockdown (Knockdown) or knockout (Knockout) sample controls. If the
band persists in the knockout sample, it is a non-specific band. |
| Tag-specific issues |
HA tag is cleaved in apoptotic cells. |
In addition to the full-length protein, there is an additional distinct small
fragment. |
Caspase cleavage site: The HA sequence contains a Caspase recognition site, which is cleaved
during apoptosis. |
| Prevention Strategy: When conducting apoptosis experiments, switch to Flag or Myc tags; or
simultaneously add tags at the C-terminus to verify the full-length protein. |
| Variability in Myc Tag Recognition |
Western Blot can detect it, but immunofluorescence (IF) or immunoprecipitation (IP)
cannot. |
Fixed-dependent epitope accessibility: The Myc epitope is masked under specific fixatives (such
as methanol) or in its native conformation. |
| Solution: In IF experiments, try replacing the permeabilization agent (e. g., Triton X-100) or
the fixative (e. g., 4% PFA). |
| Non-specific Binding of His-tag |
The background is high during detection or purification, with many impurity proteins
present. |
Metal ion contamination: Host proteins containing histidine clusters can bind to nickel columns.
|
| Buffer composition adjustment: Add low-concentration imidazole (20-40 mM) to the binding/wash
buffer to competitively elute contaminating proteins. |
9. Frequently Asked Questions (FAQ)
Q: Can I use multiple different tags on the same protein?
A: Yes. Using tandem tags (such as His-Flag, HA-Myc) can combine the advantages of
different tags, utilizing one tag for purification and another for detection or enhancing solubility.
Q: What is the smallest tag that provides reliable detection?
A: Common small tags include 6xHis (6 amino acids), Flag (8 amino acids DYKDDDDK),
and HA (9 amino acids YPYDVPDYA), all of which have well-established antibodies for use in Western
Blot and immunoprecipitation.ProductSupport.
Q: How can I determine if my tag affects protein function?
A: Validated through functional phenotypic restoration experiments.In cells with
endogenous protein knocked out, complement the expression of tagged proteins to observe whether they
can restore cellular functions (such as localization, proliferation, and signaling pathway activity)
to the same level as in wild-type cells.
Q: Which tag is most suitable for co-immunoprecipitation experiments?
A: Flag tag or Myc tag. Flag and Myc tags have low immunogenicity, fully expose
epitopes under natural conditions, and allow for mild elution conditions (e. g., Flag can be
competitively eluted using 3xFlag peptides), which helps preserve the native conformation of protein
complexes to the greatest extent.
Q: Can anti-tag antibodies cross-react with endogenous proteins?
A: Theoretically possible but rarely occurs.Tag sequences (such as YPYDVPDYA) are
extremely rare in nature, but certain endogenous proteins may contain similar epitopes. Therefore,
untransfected/unlabeled cells should be included as negative controls in every experiment.
References
[1] Tamura R, Miyazaki N, Ito Y, et al. Application of the NZ-1 Fab as a crystallization chaperone
for PA tag-inserted target proteins. Protein Sci 2019; 28: 1450–59.
[2] Fang W, Wang X, Li Y, et al. Optimized protocols for chromatin immunoprecipitation of
exogenously expressed epitope-tagged proteins. STAR Protoc 2023; 4: 102050.
[3] Hanl M, Li J, Schmidt T, et al. Target Engagement Studies and Kinetic Live-Cell Degradation
Assays Enable the Systematic Characterization of Histone Deacetylase 6 Degraders. ACS Pharmacol Transl Sci
2025; 8: 2456–68.
[4] Boursier ME, Levin S, Zimmerman K, et al. The luminescent HiBiT peptide enables selective
quantitation of G protein-coupled receptor ligand engagement and internalization in living cells. J Biol
Chem 2020; 295: 5124–35.
[5] Yoda T, Shirai Y, Okada Y, et al. Four-color single-molecule imaging system for tracking GPCR
dynamics with fluorescent HiBiT peptide. Biophys Physicobiol 2024; 21: e210020.
[6] Ankavay M, Dubuisson J, Cocquerel L, et al. Monitoring of hepatitis E virus infection and
replication by functional tagging of the ORF2 protein. JHEP Rep 2025; 7: 101293.
[7] Cabalteja CC, Mihaylov D, Cubitt B, et al. Characterization of a Nanobody-Epitope Tag
Interaction and Its Application for Receptor Engineering. ACS Chem Biol 2022; 17: 2345–56.