Hepatocellular Cancer
Hepatocellular carcinoma is the fifth most common cancer in the world and is the most common type of primary liver cancer. It is higher in people with long-term liver diseases. At present, hepatitis B virus (HBV) and hepatitis C virus (HCV) remain the most important global risk factors for HCC, but their importance will likely decline in the coming years [1]. The incidence of HCC is rising, largely attributed to a rise in hepatitis C infection.
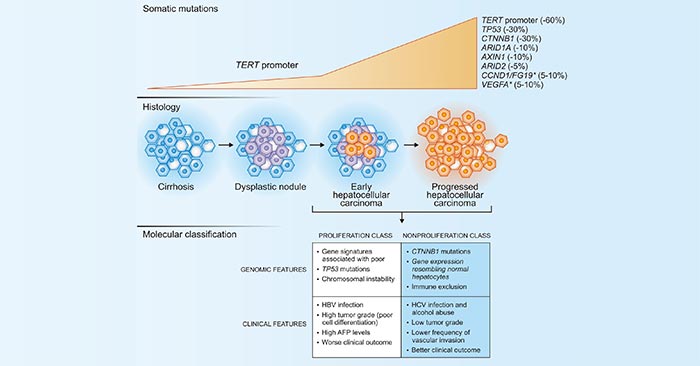
Patients with chronic liver disease have sustained hepatic inflammation, fibrosis, and aberrant hepatocyte regeneration. These abnormalities can cause cirrhosis and favor a series of genetic and epigenetic events that culminate in the formation of dysplastic nodules [2]. Hepatocellular carcinoma cells accumulate somatic DNA alterations, including mutations and chromosomal aberrations. The key molecular and histologic alterations occurring during human hepatocarcinogenesis are summarized in the figure 1. An asterisk (*) denotes high-level DNA amplification. In the nonproliferation class, immune exclusion is enriched in tumors with CTNNB1 mutations. AFP denotes alpha-fetoprotein, HBV hepatitis B virus, and HCV hepatitis C virus.
Figure 1. Main genetic alterations in hepatocellular carcinoma and molecular classification
*this diagram is derived from publication on N Engl J Med [2]
As the figure 1 shows, mutations in the TERT promoter are the most frequent genetic alterations, accounting for approximately 60% of cases [3]. They can be detected in dysplastic nodules, and the TERT promoter is a recurrent. In this article, we list part of these proteins involved in hepatocellular carcinoma based on the information provided by NCG (web resource to analyze duplicability, orthology and network properties of cancer genes).
Here, we display several key targets involved in mechanism of hepatocellular carcinoma, including:
-
TP53 (Tumor protein P53) has been described as "the guardian of the genome" because of its role in conserving stability by preventing genome mutation. Mutations in the p53 gene abrogate its normal functions, leading to genomic instability and loss of growth control. Many studies have shown that p53 is frequently mutated in HCC [4] [5] [6].
-
ARID2 (AT-rich interactive domain 2), a component of the SWItch/Sucrose Non-Fermentable (SWI/SNF) complex, functions in a chromatin remodeling complex to promote gene transcription [7]. HCC is an example of cancer that frequently harbors ARID2 mutations. Exon sequencing of 139 HCC found expected mutations in previously described genes like CTNNB1 or TP53 and unexpectedly LoF mutations in ARID2. ARID2 alterations consist in frameshift-inducing insertion or deletion, nonsense mutations, and splice-site mutations with no major hotspot of mutations [8].
-
ARID1A (AT-rich interactive domain-containing protein 1A), a component of the SWI/SNF chromatin-remodeling complex, is a recently identified tumor suppressor, s considered as a tumor suppressor gene, which is frequently mutated in hepatocellular carcinoma [9]. Chaobo Hu et al. have demonstrated ARID1A-deficiency activates Ang2-dependent angiogenesis and promotes HCC progression. Loss of ARID1A in HCCs confers sensitivity to Ang2 blockade and sorafenib treatment [10].
-
CTNNB1 (β-catenin) is a dual function protein, involved in regulation and coordination of cell–cell adhesion and gene transcription. Emerging evidence has showed that mutations in the CTNNB1 gene in hepatocellular carcinoma (HCC) are related to immune exclusion, and HCC patients with CTNNB1 mutations tend to be primarily resistant to anti-PD1 therapy [11].
-
AXIN1 (Axis inhibition protein 1), a component of the beta-catenin destruction complex, is required for regulating CTNNB1 levels through phosphorylation and ubiquitination, and modulating Wnt-signaling [12]. AXIN1 is a member of Wnt/β-catenin pathway, which is the most frequently deregulated pathway in hepatocellular carcinoma (HCC). Mutations of AXIN1 represent about 10% of HCC mutations [13].
References
[1] Katherine A McGlynn, Jessica L Petrick, Hashem B El-Serag et al. Epidemiology of Hepatocellular Carcinoma [J]. Hepatology. 2020. 10.1002.
[2] Augusto Villanueva. Hepatocellular Carcinoma [J]. N Engl J Med. 2019, 380:1450-62.
[3] Schulze K, Nault J-C, Villanueva A. Genetic profiling of hepatocellular carcinoma using next-generation sequencing [J]. J Hepatol. 2016, 65:1031-42.
[4] Jeng KS, Sheen IS, Chen BF, et al. Is the p53 gene mutation of prognostic value in hepatocellular carcinoma after resection [J]? Arch Surg. 2000, 135:1329-1333.
[5] Katiyar S, Dash BC, Thakur V et al. p53 tumor suppressor gene mutations in hepatocellular carcinoma patients in India[J]. Cancer. 2000, 88:1565-1573.
[6] George G. Chen, Juanita L. Merchant, Paul B. S. Lai, et al. Mutation of p53 in Recurrent Hepatocellular Carcinoma and Its Association with the Expression of ZBP-89 [J]. Am J Pathol. 2003. 162(6): 1823–1829.
[7] Atsushi Oba, Shu Shimada, Yoshimitsu Akiyama et al. ARID2 modulates DNA damage response in human hepatocellular carcinoma cells [J]. J Hepatol. 2017, 66(5):942-951.
[8] Robin Loesch, Linda Chenane and Sabine Colnot. Settings ARID2 Chromatin Remodeler in Hepatocellular Carcinoma [J]. Cells. 2020, 9(10), 2152.
[9] Sheng Cheng, Lan Wang, Chuan-Huai Deng et al. ARID1A represses hepatocellular carcinoma cell proliferation and migration through lncRNA MVIH [J]. Biochemical and Biophysical Research Communications. 2017, 1(491):178-182.
[10] Chaobo Hu, Weiping Li, Feng Tian et al. Arid1a regulates response to anti-angiogenic therapy in advanced hepatocellular carcinoma [J]. J Hepatol. 2018, 68(3):465-475.
[11] Xuelian Xiao, Huanye Mo, Kangsheng Tu. CTNNB1 mutation suppresses infiltration of immune cells in hepatocellular carcinoma through miRNA-mediated regulation of chemokine expression [J]. International Immunopharmacology. 2020, 89: 107043.
[12] Shuichi Kusano, Nancy Raab-Traub. I-mfa domain proteins interact with Axin and affect its regulation of the Wnt and c-Jun N-terminal kinase signaling pathways [J]. Mol Cell Biol. 2002, 22(18):6393-405.
[13] Shirley Abitbol, Rajae Dahmani, Cédric Coulouarn et al. AXIN deficiency in human and mouse hepatocytes induces hepatocellular carcinoma in the absence of β-catenin activation [J]. J Hepatol. 2018, 68(6):1203-1213.