Leukemia
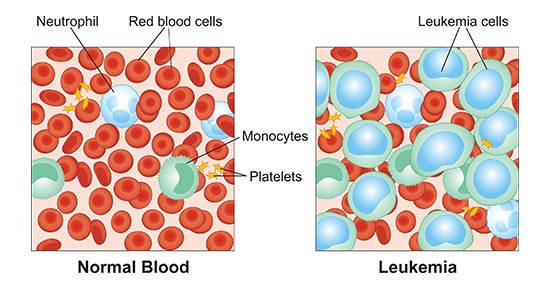
Leukemia is cancer found in the blood and bone marrow, caused by too many white blood cells in the body. White blood cells are potent infection fighters — they normally grow and divide in an orderly way, as the body needs them. But in people with leukemia, the bone marrow produces an excessive amount of abnormal white blood cells, which don't function properly (figure 1).
Figure 1. a diagram of components of normal blood and leukemia
Generally, leukemia can be divided into four types, including acute lymphocytic leukemia (ALL), acute myeloid leukemia (AML), chronic lymphocytic leukemia (CLL) and chronic myeloid leukemia (CML). Among of them, AML is the most common type of acute leukemia in adults. It occurs in children and adults.
Factors that may increase your risk of developing some types of leukemia include previous cancer treatment, genetic disorders, exposure to certain chemicals, smoking and family history of leukemia. Among of them, genetic abnormalities seem to play a role in the development of leukemia. Certain genetic disorders, such as Down syndrome, are associated with an increased risk of leukemia.
In this article, we list part of these proteins involved in leukemia based on the information provided by NCG (web resource to analyze duplicability, orthology and network properties of cancer genes).
Here, we display several key targets involved in mechanism of leukemia, including:
-
NRAS (neuroblastoma RAS) is a member of RAS (rat sarcoma) family, which also includes KRAS (Kirsten RAS), and HRAS (Harvey RAS). Ras proteins comprise a family of low-molecular-weight GTPases. Wild-type RAS serves a critical role in cellular proliferation [1]. NRAS mutations are present in 15% to 20% of melanomas, AMLs (10%) and 8% to 10% of thyroid cancers [2]. Moreover, NRAS mutations are also present in a variety of other hematologic malignancies, including ALL (11%), multiple myeloma (18%), MDS (5%), and CML (19%) [3] [4].
-
DNMT3A (NA (cytosine-5)-methyltransferase 3A) is an enzyme that catalyzes the transfer of methyl groups to specific CpG structures in DNA, a process called DNA methylation. The study executed by Hsin-An Hou et al. reported that DNMT3A mutations were identified in 14% of total patients and in 22.9% of AML patients with normal karyotype in the present study of 500 patients with de novo AML. DNMT3A mutations were positively associated with older age, higher WBC and platelet counts, intermediate-risk and normal cytogenetics, FLT3 internal tandem duplication, and NPM1, PTPN11, and IDH2 mutations, but were negatively associated with CEBPA mutations [5].
-
RUNX1 (Runt-related transcription factor 1), also known as acute myeloid leukemia 1 protein, binds to specific regions of DNA and helps control the activity of particular genes. It is widely expressed in hematopoietic cells. Many lines of evidence have shown that RUNX1 plays a critical role in regulating the development and precise maintenance of mammalian hematopoiesis [6]. RUNX1 is generally considered a tumor suppressor in myeloid neoplasms. Inactivating RUNX1 mutations have frequently been found in patients with myelodysplastic syndrome (MDS) and cytogenetically normal AML [7].
-
TET2 (ten-eleven translocation 2), is involved in regulating the process of transcription. This protein is a tumor supressor and mutations of the gene are seen in myeloid malignancies and other hematological disorders. Mutations and deletions were detected in myelodysplastic syndromes, acute myeloid leukemias (AML) and other myeloid malignancies [8] [9].
-
FLT3 (FMS-like tyrosine kinase 3) is a cytokine receptor which belongs to the receptor tyrosine kinase class III. It plays a critical roles in in hematopoietic stem/progenitor cell survival and proliferation. It is mutated in about 1/3 of acute myeloid leukemia (AML) patients, either by internal tandem duplications (ITD) of the juxtamembrane domain or by point mutations usually involving the kinase domain (KD) [10].
-
NOTCH1 (Notch homolog 1), functions as a receptor for membrane-bound ligands Jagged-1 (JAG1), Jagged-2 (JAG2) and Delta-1 (DLL1) to regulate cell-fate determination. Accumulating evidence indicates a key role of deregulated NOTCH1 signaling and NOTCH1 mutations in CLL [11]. Chromosomal rearrangements and mutations of the NOTCH1 gene were initially described in T-cell acute lymphoblastic leukemia (T-ALL), which displayed aberrant activation of NOTCH1 signaling in over 60% of the cases [12].
References
[1] Douglas B. Johnson, Keiran S.M. Smalley and Jeffrey A. Sosman. Molecular Pathways: Targeting NRAS in Melanoma and Acute Myelogenous Leukemia [J]. Clin Cancer Res. 2014, 20 (16) : 4186-4192.
[2] Pylayeva-Gupta Y, Grabocka E, Bar-Sagi D. RAS oncogenes: weaving a tumorigenic web [J]. Nat Rev Cancer. 2011, 11:761–74.
[3] Ward AF, Braun BS, Shannon KM. Targeting oncogenic Ras signaling in hematologic malignancies [J]. Blood. 2012, 120:3397–406.
[4] Irahara N, Baba Y, Nosho K et al. NRAS mutations are rare in colorectal cancer [J]. Diagn Mol Pathol. 2010, 19:157–63.
[5] Hsin-An Hou, Yuan-Yeh Kuo, Chieh-Yu Liu et al. DNMT3A mutations in acute myeloid leukemia: stability during disease evolution and clinical implications [J]. Blood. 2012, 119 (2): 559–568.
[6] Ichikawa M, Yoshimi A, Nakagawa M et al. A role for RUNX1 in hematopoiesis and myeloid leukemia [J]. Int J Hematol. 2013, 97 (6):726-34.
[7] Goyama S, Schibler J, Cunningham L et al. Transcription factor RUNX1 promotes survival of acute myeloid leukemia cells [J]. J Clin Invest. 2013, 123 (9): 3876-88.
[8] Tefferi A, Pardanani A, Lim KH et al. TET2 mutations and their clinical correlates in polycythemia vera, essential thrombocythemia and myelofibrosis [J]. Leukemia. 2009, 23: 905–911.
[9] Jankowska AM, Szpurka H, Tiu RV et al. Loss of heterozygosity 4q24 and TET2 mutations associated with myelodysplastic/myeloproliferative neoplasms [J]. Blood. 2009, 113: 6403–6410.
[10] Small D. FLT3 mutations: biology and treatment [J]. Hematology Am Soc Hematol Educ Program. 2006:178-84.
[11] Rosati E, Baldoni S, De Falco F et al. NOTCH1 Aberrations in Chronic Lymphocytic Leukemia [J]. Front Oncol. 2018, 8:229.
[12] Emanuela Rosati, Stefano Baldoni, Filomena De Falco et al. NOTCH1 Aberrations in Chronic
Lymphocytic Leukemia [J]. Front. Oncol. 2018, 8:229.