Ovarian Cancer
Ovarian cancer is a type of cancer that begins in the ovaries, which produce eggs (ova) as well as the hormones estrogen and progesterone. Ovarian cancer often goes undetected until it has spread within the pelvis and abdomen. Ovarian cancers can be classified into three large groups, including epithelial, germ cell, and specialized stromal cell tumors. Among of them, the largest majority of ovarian cancers are epithelial ovarian cancers (EOCs). EOC can be further divided into two main histological subtypes: Type I and Type II tumors. Type I tumors tend to grow more slowly, often from an identifiable precursor. However, type II tumors are characterized by high-grade and rapidly progressive disease [1].
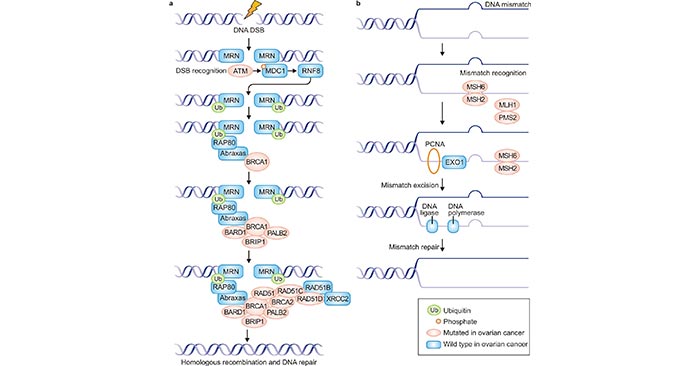
Several factors can increase the risk of developing ovarian cancer, including genetic factors, age, postmenopausal hormonal therapy use, infertility and nulliparity [2]. Currently, the best-studied genetic alterations in ovarian cancers are those involved in DNA repair (Figure 1). As the figure 1a shows, the double-stranded DNA break and homologous repair process begins with recognition and sensing of double-strand breaks (DSBs) by the meiotic recombination 11 homologue 1 (MRE11)–RAD50–Nijmegen breakage syndrome protein 1 (NBS1) (MRN) complex, which acts as an activation site for the serine-protein kinase ATM. In the figure 1b, DNA mismatch repair is mediated by the MutS protein homologue 2 (MSH) proteins, as well as the endonuclease PMS2 and proliferating cell nuclear antigen (PCNA).
Figure 1. DNA repair mechanisms and ovarian cancer
*this diagram is derived from publication on Nat Rev Dis Primers [2]
Several studies have revealed that the commonly implicated inherited genes in ovarian cancer are BRCA1, BRCA2 and BRIP1, genes that are part of the Fanconi anaemia pathway (RAD51C, RAD51D, BRIP1,PALB2 and BARD1) and genes that are involved in DNA mismatch repair (MSH2, MSH6, MLH1 and PMS2) [3] [4] [5] [6]. In this article, we list part of targets involved in ovarian cancer based on the information provided by NCG (web resource to analyze duplicability, orthology and network properties of cancer genes).
Here, we display several key targets involved in mechanism of ovarian cancer, including:
-
NF1 (Neurofibromatosis type 1), a RAS GTPase-activating protein, which negatively regulates Ras signaling [7]. Emerging study implicated a link between neurofibromatosis type 1 and ovarian cancer [8]. This study found that biallelic inactivation of the NF1 gene is an early event in high grade serous ovarian cancer (HGSOC) tumorigenesis. The NF1 pathway is a potential therapeutic target in women with NF1 that develop HGSOC.
-
BRCA1 (Breast cancer type 1 susceptibility protein) and BRCA2 (Breast cancer type 2 susceptibility protein) are crucially involved in the repair of double-strand breaks by homologous recombination. They serve as a scaffold for other proteins involved in double-strand DNA repair, mostly through defective homologous recombination [9]. They can stabilizes RAD51–ssDNA complexes. Germline mutations in BRCA1 are present in about 5% of women with ovarian cancer.
-
KRAS (Kirsten rat sarcoma viral oncogene homolog) is a member of MAP kinase (MAPK) pathway. Mutations in the KRAS gene are one of the most frequent genetic abnormalities in ovarian carcinoma. They are of renewed interest as new epidermal growth factor receptor (EGFR)-targeted therapies are being investigated for use in ovarian carcinoma [10].
-
RB1 (retinoblastoma, pRb), a tumor suppressor protein, is dysfunctional in several major cancers. The function of RB1 is to prevent excessive cell growth by inhibiting cell cycle progression until a cell is ready to divide. pRb is a negative regulator of cellular proliferation, which is achieved by sequestering a variety of nuclear proteins involved in cellular growth. Alterations in the pRb pathway are observed frequently in epithelial ovarian cancer (EOC); ∼30% of ovarian cancers exhibit loss of heterozygosity at the RB1 gene locus [11].
References
[1] Paul T. Kroeger Jr. and Ronny Drapkin. Pathogenesis and heterogeneity of ovarian cancer [J]. Curr Opin Obstet Gynecol. 2017, 29:26–34.
[2] Ursula A. Matulonis, Anil K. Sood, Lesley Fallowfield et al. Ovarian cancer [J]. Nat Rev Dis Primers. 2016, 2:16061.
[3] Pennington, K. P. & Swisher, E. M. Hereditary ovarian cancer: beyond the usual suspects [J]. Gynecol. Oncol. 2012, 124, 347–353.
[4] Walsh, T. et al. Mutations in 12 genes for inherited ovarian, fallopian tube, and peritoneal carcinoma identified by massively parallel sequencing [J]. Proc. Natl Acad. Sci. USA. 2011, 108, 18032–18037.
[5] Norquist, B. M. et al. Inherited mutations in women with ovarian carcinoma [J]. JAMA Oncol. 2016, 2, 482–490.
[6] Prakash, R., Zhang, Y., Feng, W. & Jasin, M. Homologous recombination and human health: the roles of BRCA1, BRCA2, and associated proteins [J]. Cold Spring Harb. Perspect. Biol. 2015, 7, a016600.
[7] Eliza Courtney, Sock Hoai Chan, Shao Tzu Li et al. Biallelic NF1 inactivation in high grade serous ovarian cancers from patients with neurofibromatosis type 1 [J]. Fam Cancer. 2020, 19(4):353-358.
[8] Eric J.Norris, Wendell D.Jones, Marius D.Surleac et al. Clonal lineage of high grade serous ovarian cancer in a patient with neurofibromatosis type 1 [J]. Gynecologic Oncology Reports. 2018, 23: 41-44.
[9] O’Donovan, P. J. & Livingston, D. M. BRCA1 and BRCA2: breast/ovarian cancer susceptibility gene products and participants in DNA double-strand break repair [J]. Carcinogenesis. 2010, 31, 961–967.
[10] Veronika Auner, Gernot Kriegshäuser, Dan Tong et al. KRAS mutation analysis in ovarian samples using a high sensitivity biochip assay [J]. BMC Cancer. 2009, 9, 111.
[11] Honglin Song, Susan J. Ramus, Danielle Shadforth et al. Common Variants in RB1 Gene and Risk of Invasive Ovarian Cancer [J]. Cancer Research. 2006.