Bladder Cancer
Bladder cancer is a major malignant tumor with a high incidence in the urinary system that begins in the cells of the bladder. The bladder is a hollow muscular organ in your lower abdomen that stores urine. Based on the types of bladder cells where cancer begins, the types of bladder cancer can be divided into three groups, including urothelial carcinoma, squamous cell carcinoma and adenocarcinoma. Urothelial cells is the cells that line the inside of your bladder. Bladder cancer most often begins in these cells.
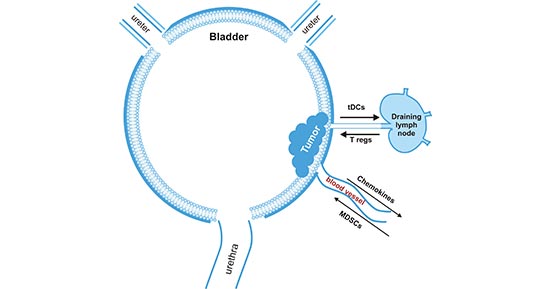
Bladder cancer represents an ideal disease state to study immune evasion and mechanisms by which to improve the immune response based on several established features [1]. One of the hallmarks of cancer progression is a formation of immunosuppressive and tolerogenic tumor microenvironment (figure 1) [2].
Figure 1. Bladder tumor-induced immune suppression promotes the escape of malignant epithelial bladder cells from the immune system
*this diagram is derived from publication on Cancer Immunology [2]
Additionally, the gene alternations also are important in the progress of bladder cancer. Lucie C Kompier et al. found that of primary tumors 64% were mutant for FGFR3, 11% for RAS, 24% for PIK3CA, and 26% for p53 by mutation assays [3]. In this article, we list part of these proteins involved in bladder cancer based on the information provided by NCG (web resource to analyze duplicability, orthology and network properties of cancer genes).
Here, we display several key targets involved in mechanism of bladder cancer, including:
-
STAG2 (Stromal Antigen 2), one of four components of the cohesin complex, mediates cohesion between sister chromatids following DNA replication to ensure correct chromosomal segregation [4]. STAG2 is the most frequently mutated cohesin subunit and was recently identified as a gene that is commonly altered in bladder cancer [5]. Alana Lelo et al. found that STAG2 mutations were much more common in non–muscle-invasive bladder cancer (NMIBC) (∼32%) than in tumors that had progressed to muscle invasion [6] [7].
-
EP300 (E1A binding protein P300), also called P300, functions as histone acetyltransferase which regulates transcription of genes via chromatin remodeling by allowing histone proteins to wrap DNA less tightly. This protein plays an essential role in regulating cell growth and division, prompting cells to mature and take on specialized functions. EP300 is frequently mutated in bladder cancer, and its mutation is associated with increased TMB and promotes antitumor immunity, which may serve as a biomarker to predict immune response [8].
-
FGFR3 (fibroblast growth factor receptor 3), also known as CD333, plays an essential role in the regulation of cell proliferation, differentiation and apoptosis. Alterations affecting FGFR3 signaling are found more frequently in bladder than in any other cancer type. Aberrant activation of the receptor can occur via several mechanisms. These are related to tumor stage and grade, and have different effects on downstream signalling and phenotypic consequences [9].
-
KDM6A (Lysine (K)-specific demethylase 6A) is found in many organs and tissues of the body and functions in the catalysis of the demethylation of tri/dimethylated histone H3. It is a histone modifier frequently mutated in bladder cancer. The study of Kohei Kobatake et al. suggested that Kdm6a deficiency activated cytokine and chemokine pathways, promoted M2 macrophage polarization, increased cancer stem cells and caused bladder cancer in cooperation with p53 haploinsufficiency [10].
-
ARID1A (AT-rich interactive domain-containing protein 1A), a part of a protein complex called SWI/SNF, which regulate gene activity (expression) by a process known as chromatin remodeling. SWI/SNF complexes are mutated in ~20% across human cancers. And Stefan Garczyk et al. reported that ARID1A truncating mutations, potentially translating into ARID1A protein loss in a subset of high-grade bladder cancers, are the most common SWI/SNF genetic alterations in bladder cancer [11].
References
[1] Alegrezza MJ, Conejo-Garcia JR. Targeted therapy and immunosuppression in the tumor microenvironment [J]. Trends Cancer. 2017, 3(1):19–27.
[2] Paul L. Crispen & Sergei Kusmartsev. Mechanisms of immune evasion in bladder cancer [J]. Cancer Immunology. 2020, 3–14.
[3] Lucie C Kompier, Irene Lurkin, Madelon N M van der Aa et al. FGFR3, HRAS, KRAS, NRAS and PIK3CA mutations in bladder cancer and their potential as biomarkers for surveillance and therapy [J]. PLoS One. 2010, 5(11):e13821.
[4] Claire F. Taylor, Fiona M. Platt and Carolyn D. Hurst. Frequent inactivating mutations of STAG2 in bladder cancer are associated with low tumour grade and stage and inversely related to chromosomal copy number changes [J]. Hum Mol Genet. 2014, 23(8): 1964–1974.
[5] Lanni Aquila, Joyce Ohm and Anna Woloszynska. The role of STAG2 in bladder cancer [J]. Pharmacological Research. 2019, 142:143-149.
[6] Solomon DA, Kim JS, Bondaruk J et al. Frequent truncating mutations of STAG2 in bladder cancer [J]. Nat Genet. 2013, 45:1428–30.
[7] Alana Lelo, Frederik Prip, Brent T. Harris et al. STAG2 Is a Biomarker for Prediction of Recurrence and Progression in Papillary Non–Muscle-Invasive Bladder Cancer [J]. Clin Cancer Res. 2018, (24) (17):4145-4153.
[8] Gongmin Zhu, Lijiao Pei, Yuan Li et al. EP300 mutation is associated with tumor mutation burden and promotes antitumor immunity in bladder cancer patients [J]. Aging (Albany NY). 2020, 12(3):2132-2141.
[9] Knowles and Margaret A. FGFR3 – a Central Player in Bladder Cancer Pathogenesis [J]? Bladder Cancer. 2020, 6(4):403-423.
[10] Kohei Kobatake, Ken-ichiro Ikeda, Yuichiro Nakata et al. Kdm6a Deficiency Activates Inflammatory Pathways, Promotes M2 Macrophage Polarization, and Causes Bladder Cancer in Cooperation with p53 Dysfunction [J]. Clin Cancer Res. 2020, 26:2065–79.
[11] Garczyk S, Schneider U, Lurje I et al. ARID1A-deficiency in urothelial bladder cancer: No predictive biomarker for EZH2-inhibitor treatment response [J]? 2018, PLoS ONE. 13(8): e0202965.