Phosphatidylinositol 3-kinase (PI3K) is a lipid kinase that plays essential roles in cellular signaling pathways regulating cell growth, proliferation, survival, and metabolism. This enzyme phosphorylates the 3-hydroxyl group of phosphatidylinositol and its phosphorylated derivatives, generating important second messengers that activate downstream effectors including AKT/PKB. PI3K signaling is frequently dysregulated in various diseases, particularly cancer. This makes it an important target for therapeutic intervention and biomedical research.
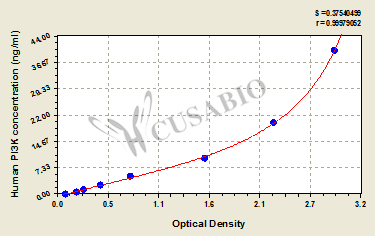
The Human Phosphotylinosital 3 kinase,PI3K ELISA Kit (CSB-E08417h) provides quantitative measurement of PI3K in human samples using a sandwich assay principle. This kit works with serum, tissue homogenates, and cell lysates. It has a detection range of 0.625 ng/mL to 40 ng/mL and sensitivity of 0.156 ng/mL. The assay requires 50-100 μL sample volume, measures absorbance at 450 nm wavelength, and can be completed within 1-5 hours.
Application Examples
Note: The following application examples are drawn from a selection of publications citing this product. For additional applications, please refer to the full list of references in the "Citations" section.
This ELISA kit has been used in research examining cellular signaling pathways and protein quantification studies. The kit allows measurement of PI3K activity in various experimental contexts, supporting studies of molecular mechanisms and biomarker analysis.
• Cellular signaling research: Used to quantify PI3K levels alongside other key signaling proteins including EGFR, Akt, and PDK1 in pathway analysis studies
• Protein expression profiling: Applied in comparative studies examining protein levels across different experimental treatment groups
• Biomarker research: Used in studies measuring PI3K activity in conjunction with amyloid-beta protein analysis
• Molecular mechanism studies: Applied to measure PI3K levels as part of broader studies into cellular processes and protein interactions