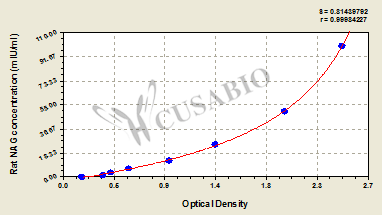
N-acetyl-β-D-glucosaminidase (NAG) is a lysosomal enzyme that breaks down glycosaminoglycans and glycoproteins within cellular lysosomes. This hexosaminidase catalyzes the hydrolysis of terminal N-acetylglucosamine residues from various substrates and serves as an important biomarker for lysosomal function and cellular damage. Elevated NAG levels in biological fluids, particularly urine, indicate renal tubular damage, making it a valuable diagnostic and research tool in nephrology and toxicology studies.
The Rat N-acetyl-β-D-glucosaminidase,NAG ELISA Kit (CSB-E07443r) is a quantitative sandwich immunoassay for measuring NAG levels in rat samples. This kit works with multiple sample types including serum, plasma, cell culture supernates, tissue homogenates, and urine, with a detection range of 1.56-100 mIU/mL and sensitivity of 0.39 mIU/mL. The assay requires 50-100 μL sample volume, operates with detection at 450 nm wavelength, and can be completed within 1-5 hours.
Application Examples
Note: The following application examples are drawn from a selection of publications citing this product. For additional applications, please refer to the full list of references in the "Citations" section.
This ELISA kit has been used in renal function research to measure kidney damage and dysfunction through urinary biomarker analysis. The kit provides a valuable tool for evaluating tubular injury in experimental studies investigating nephrotoxicity and kidney pathology.
• Renal toxicity studies - Measurement of kidney damage in experimental models examining drug-induced nephrotoxicity and protective interventions
• Urinary biomarker analysis - Quantification of tubular injury markers in urine samples as indicators of renal dysfunction
• Kidney function evaluation - Integration with comprehensive renal panels including traditional markers of kidney health